Research Articles
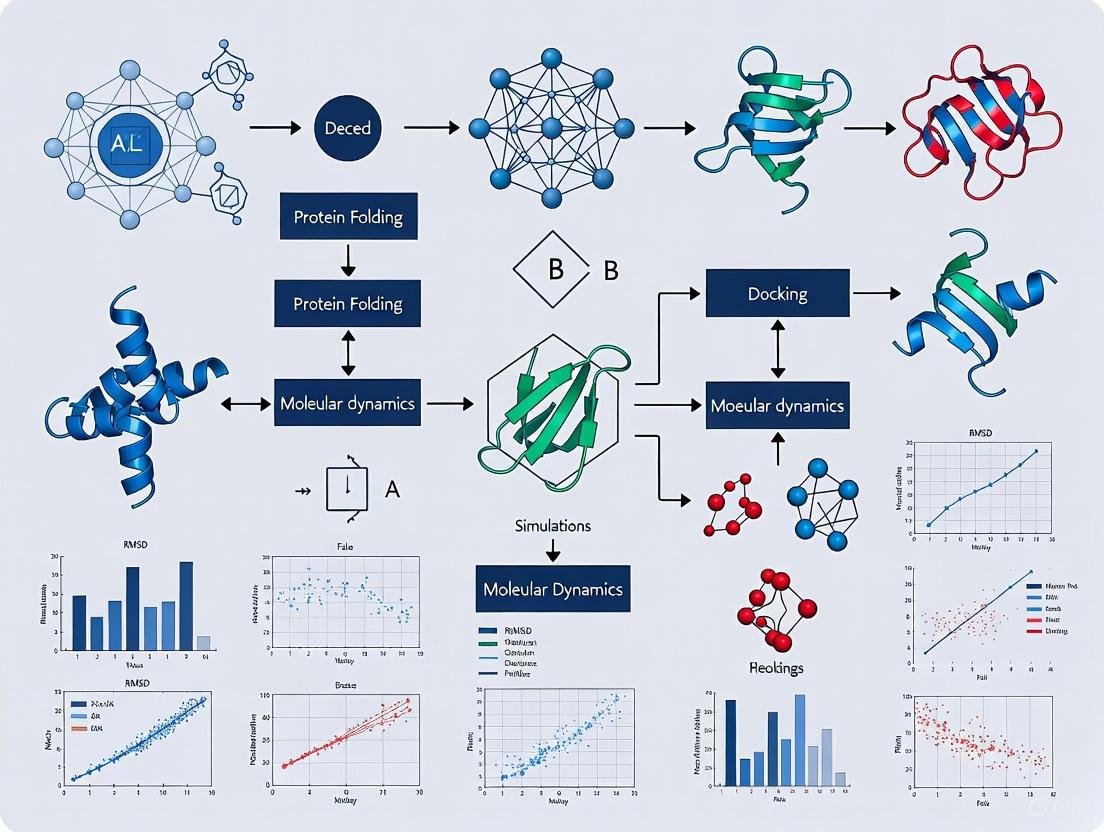
Deep Learning for Protein Representation Encoding: From Sequence to Structure in Computational Biology
Protein representation learning (PRL) has emerged as a transformative force in computational biology, enabling data-driven insights into protein structure and function.
From Sequence to Therapy: Decoding Protein Structure-Function Relationships for Advanced Research and Drug Development
This article provides a comprehensive analysis of the protein sequence-structure-function relationship, a cornerstone of molecular biology with critical implications for biomedical research and therapeutic discovery.
Protein Language Models: How Transformer Architectures Are Revolutionizing Drug Discovery and Bioinformatics
This article provides a comprehensive overview of Protein Language Models (PLMs), deep learning systems based on Transformer architectures that are transforming computational biology and drug discovery.
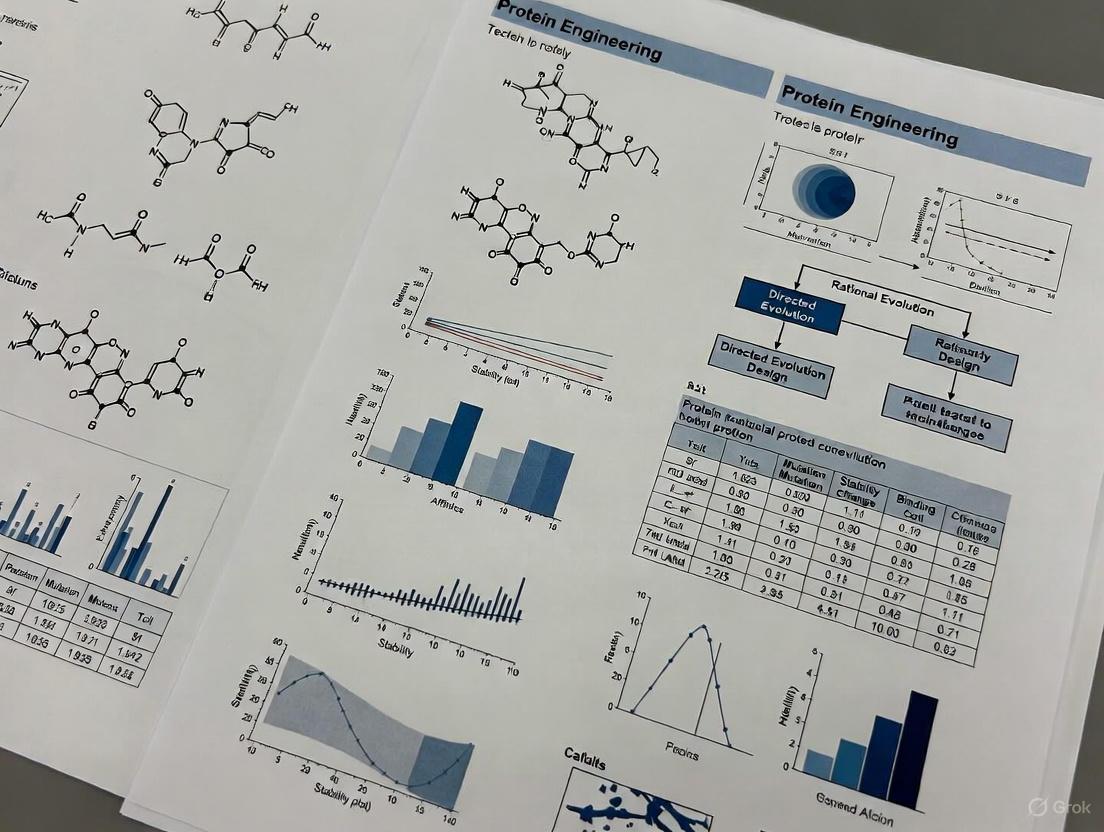
Marginal Stability in Proteins: An Evolutionary Quirk Turned Engineering Challenge
This article explores the pervasive phenomenon of marginal protein stability, examining its origins in evolutionary dynamics and its profound implications for modern protein engineering and therapeutic development.
Geometric Deep Learning for Protein Structures: A New Paradigm in Drug Discovery and Protein Design
This article provides a comprehensive exploration of geometric deep learning (GDL) and its transformative impact on computational biology, specifically for analyzing and designing protein structures.
Preventing Machine Learning Overfitting on Protein Data: Strategies for Robust Models in Drug Discovery
Overfitting presents a critical challenge in applying machine learning to protein science, where complex models can memorize noise and dataset-specific artifacts instead of learning generalizable biological principles.
Zero-Shot Prediction Revolutionizes Protein Engineering: From AI Models to Real-World Applications
This article explores the transformative impact of zero-shot AI models on protein engineering, a paradigm that predicts the functional effects of protein mutations without requiring task-specific training data.
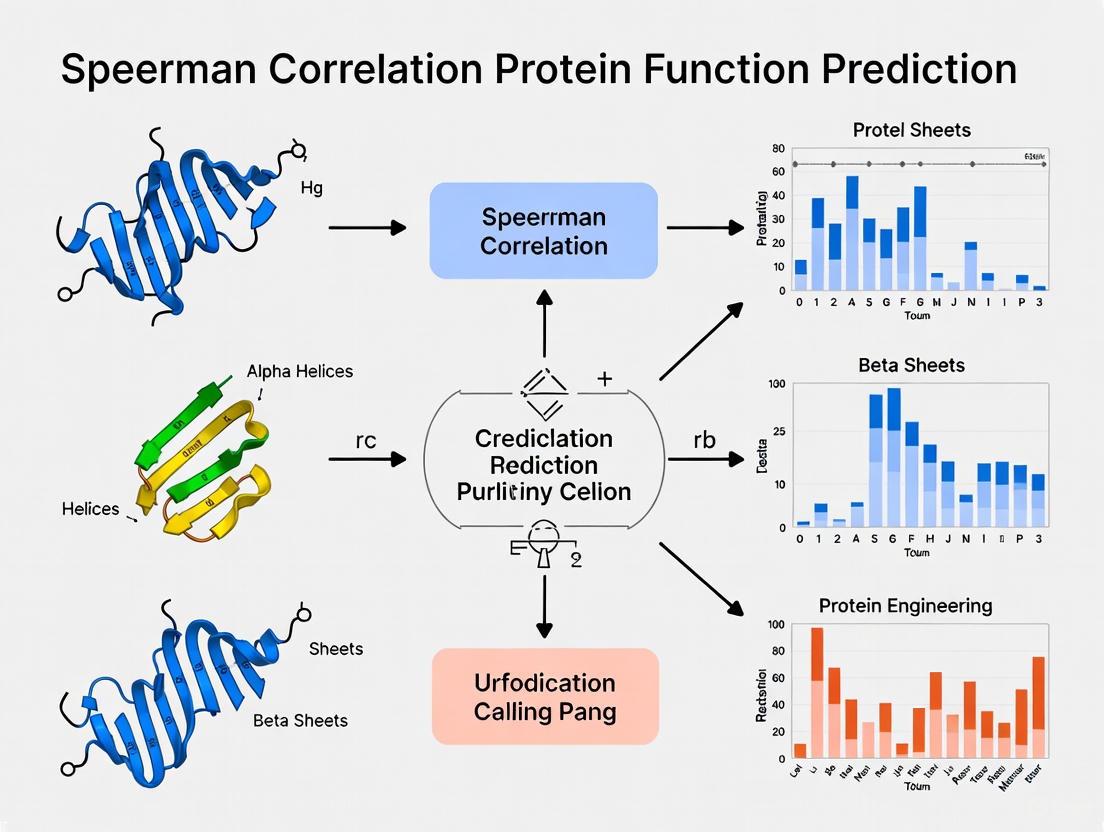
Beyond Pearson: Why Spearman Correlation is Powering the Next Generation of Protein Function Prediction
This article explores the critical role of Spearman's rank correlation coefficient in advancing the field of protein function prediction.
From Silicon to Lab Bench: A Modern Guide to Validating Computational Protein Designs
This article provides a comprehensive roadmap for researchers and drug development professionals navigating the critical process of experimentally validating computational protein designs.
Comparative Assessment of Protein Large Language Models: Performance, Applications, and Future Directions in Bioinformatics and Drug Discovery
Protein Large Language Models (PLMs), built on transformer architectures, are revolutionizing the analysis of protein sequences for function prediction, structure determination, and de novo design.