Research Articles
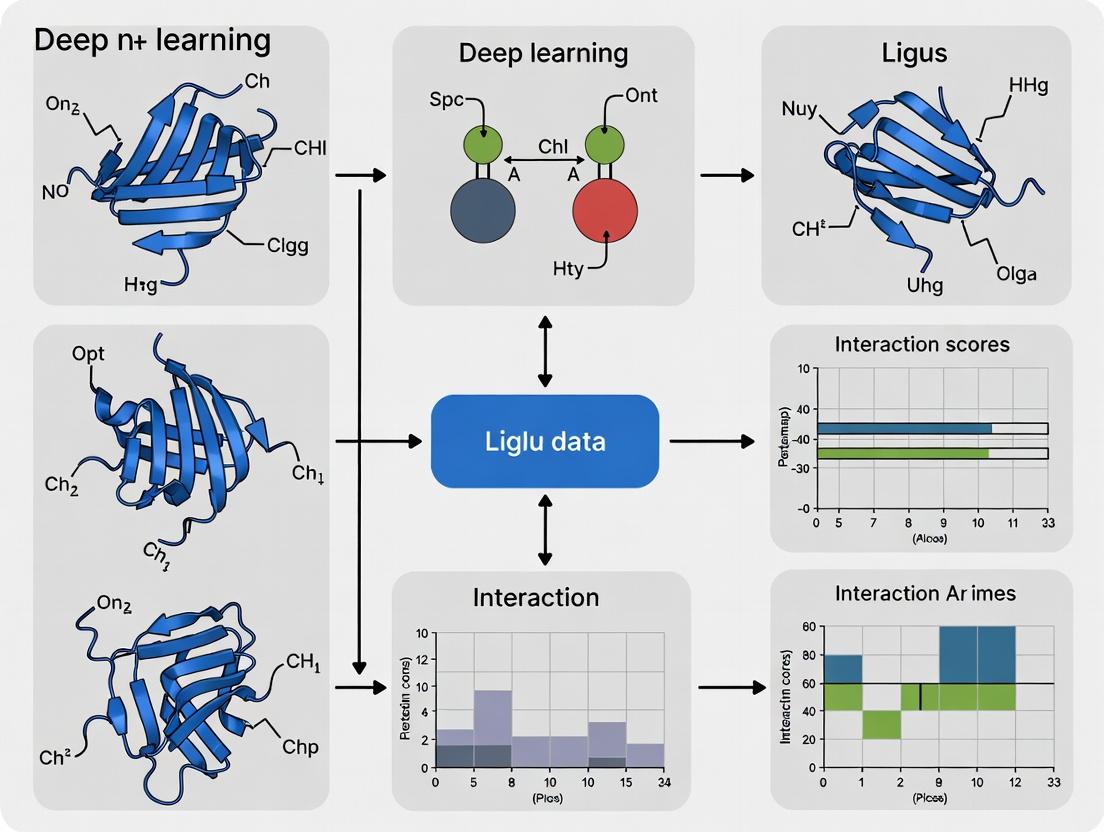
Beyond Docking: How Deep Learning Revolutionizes Protein-Ligand Interaction Prediction in Drug Discovery
This article provides a comprehensive guide for researchers and drug development professionals on the transformative role of deep learning (DL) in predicting protein-ligand interactions (PLI).
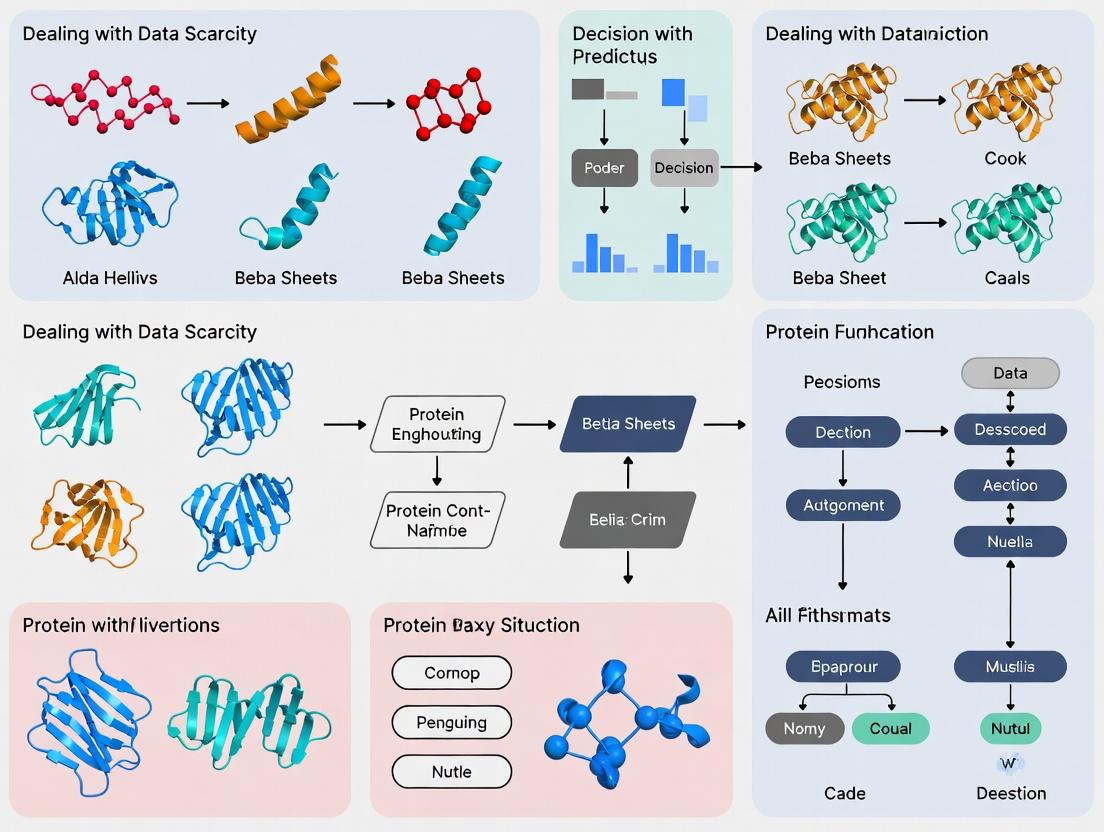
Protein Function Prediction with Limited Data: Innovative Strategies to Overcome Data Scarcity in Biomedical AI
This article addresses the critical challenge of data scarcity in protein function prediction, a major bottleneck in computational biology and AI-driven drug discovery.
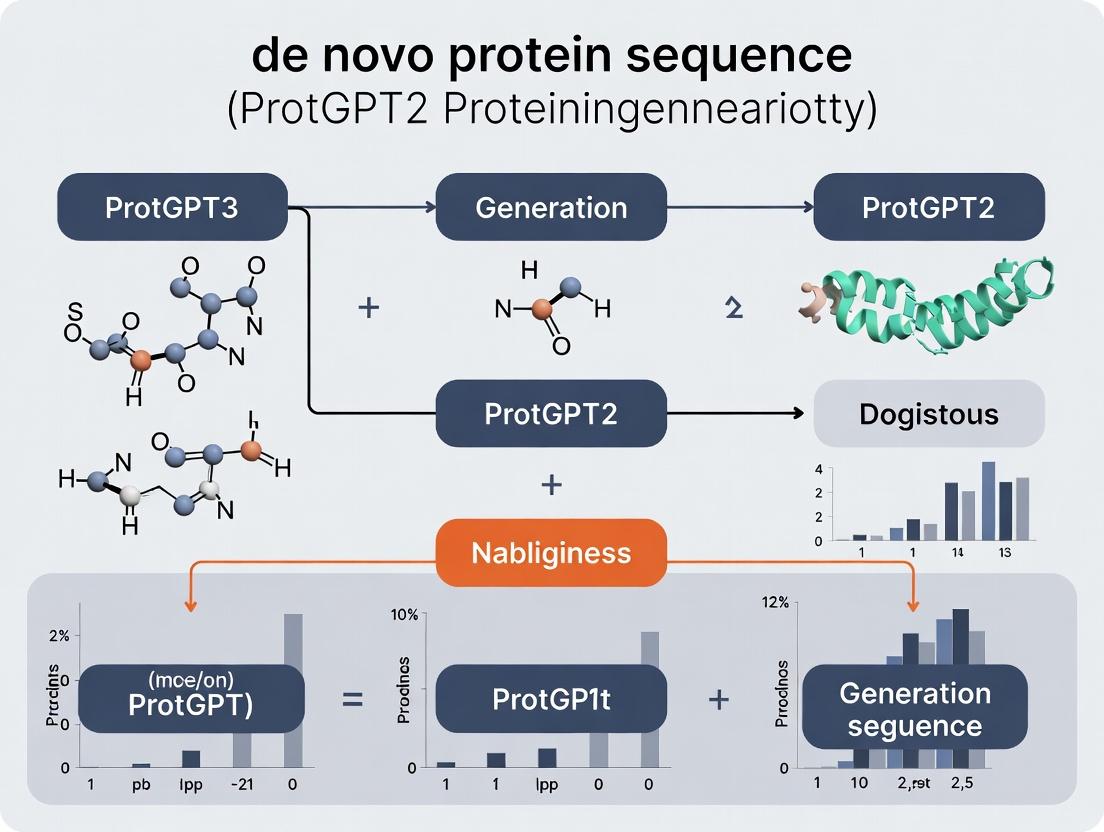
ProtGPT2: A Practical Guide to De Novo Protein Sequence Generation for Drug Discovery and Protein Design
This article provides a comprehensive guide to ProtGPT2, a transformer-based language model for generating novel, stable protein sequences.
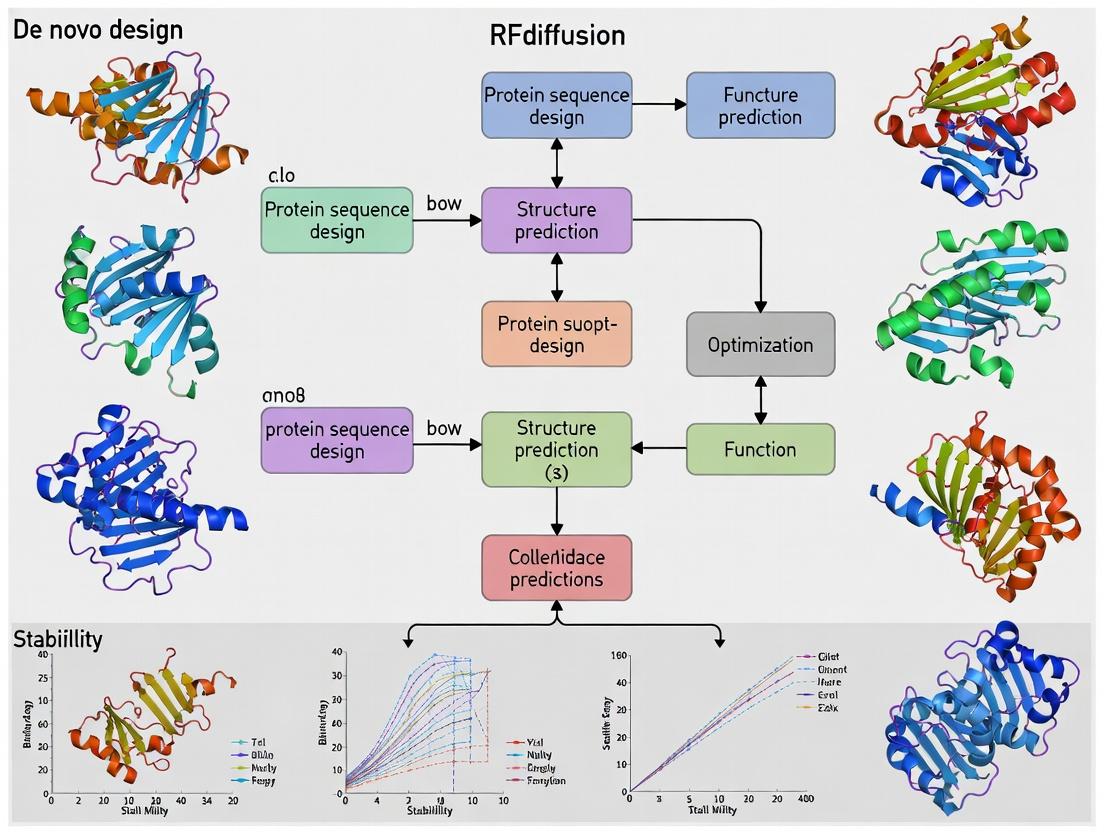
RFdiffusion: The AI-Powered Revolution in De Novo Protein Design for Therapeutics and Research
This article provides a comprehensive guide to RFdiffusion, a groundbreaking deep learning model for designing novel protein structures and functions from scratch.
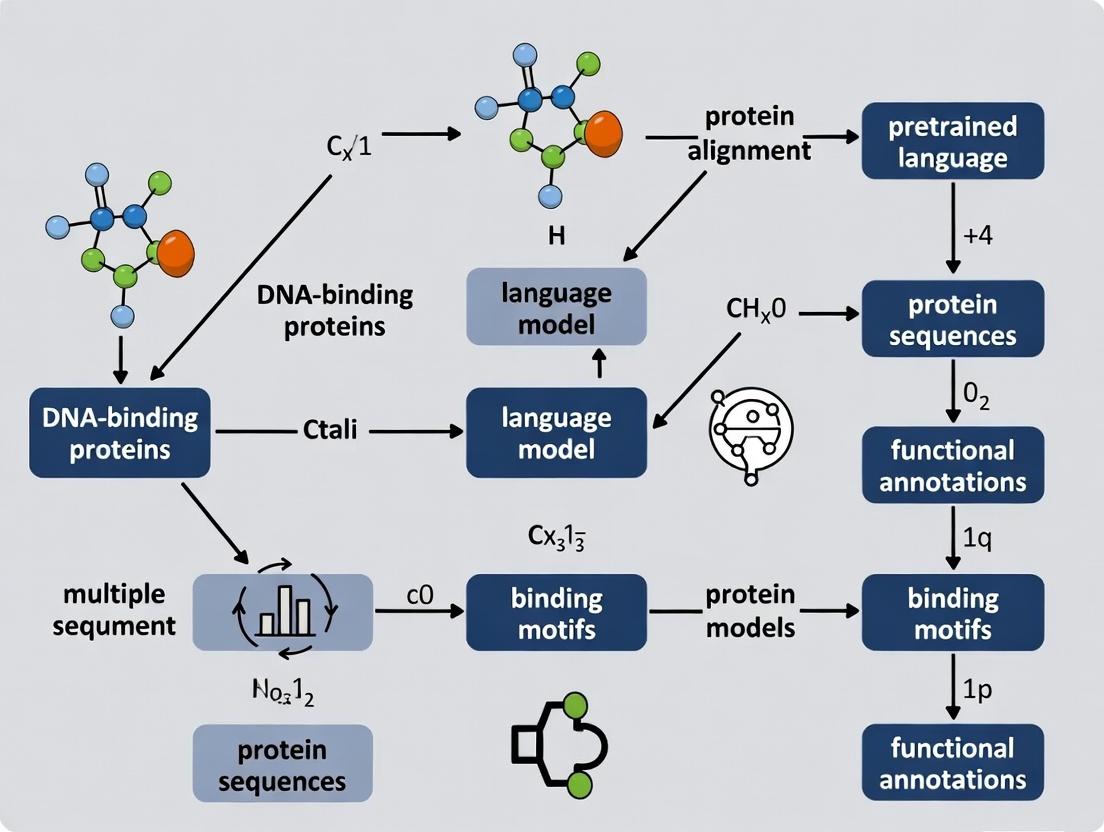
ProteinDNABERT and Beyond: How Pretrained Language Models Are Revolutionizing DNA-Binding Protein Identification
This article provides a comprehensive guide for researchers and drug discovery scientists on the application of pretrained language models (PLMs) like ProtBERT, ESM, and ProteinDNABERT for identifying DNA-binding proteins.
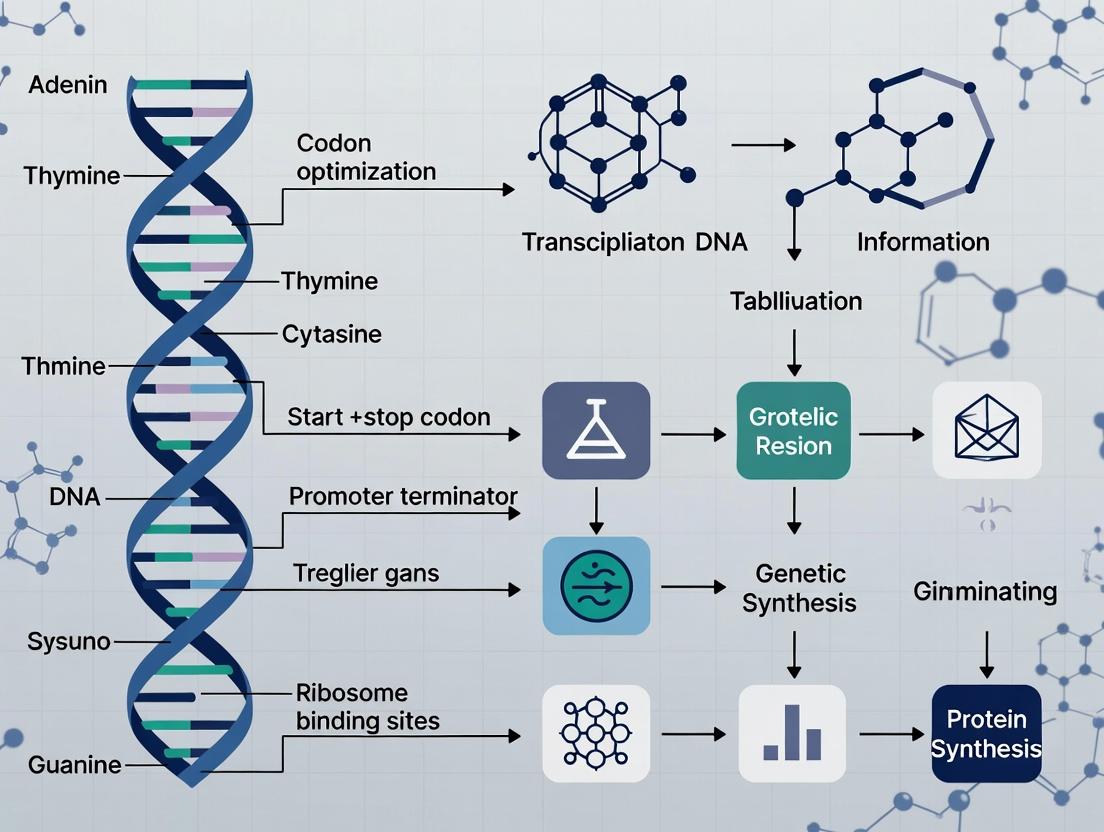
Codon Optimization for CFPS: A Complete Guide to DNA Template Design for High-Yield Protein Synthesis
This comprehensive guide explores the critical role of DNA template design and codon optimization in Cell-Free Protein Synthesis (CFPS) systems.
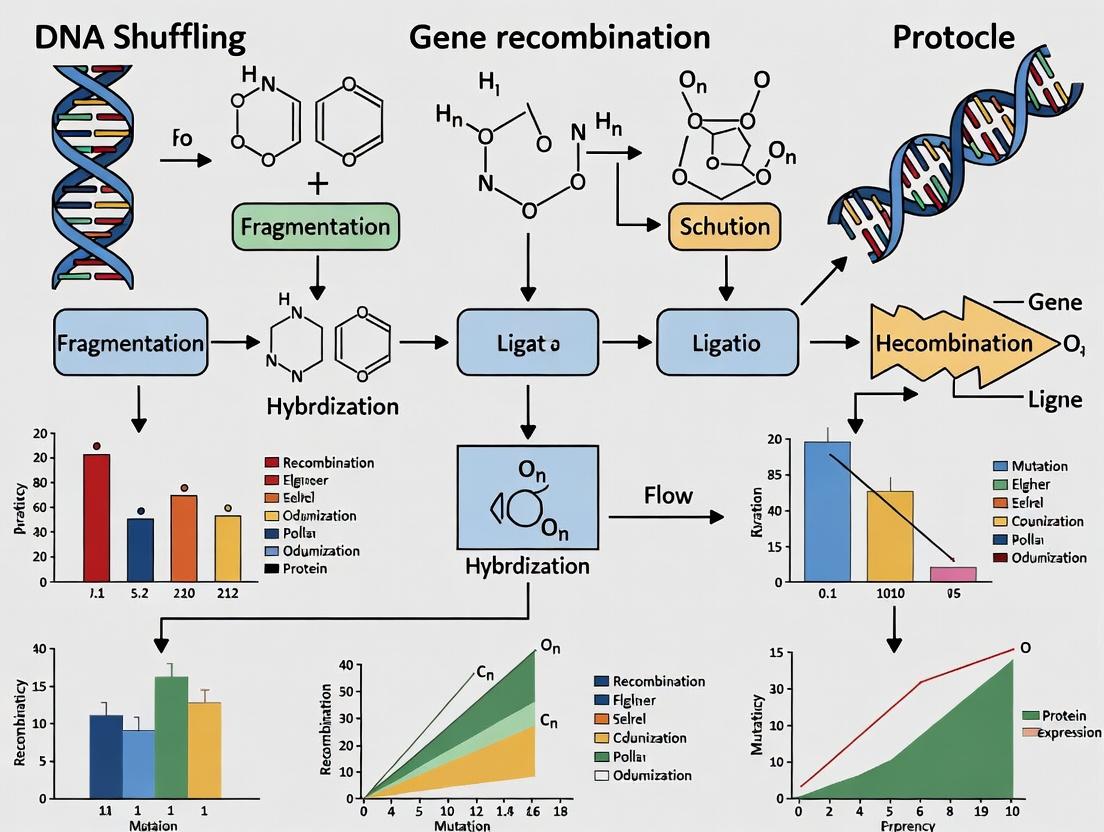
Advanced DNA Shuffling and Gene Recombination Protocols: A Comprehensive Guide for Accelerating Protein Engineering and Drug Discovery
This comprehensive guide details modern DNA shuffling and gene recombination protocols for researchers and drug development professionals.
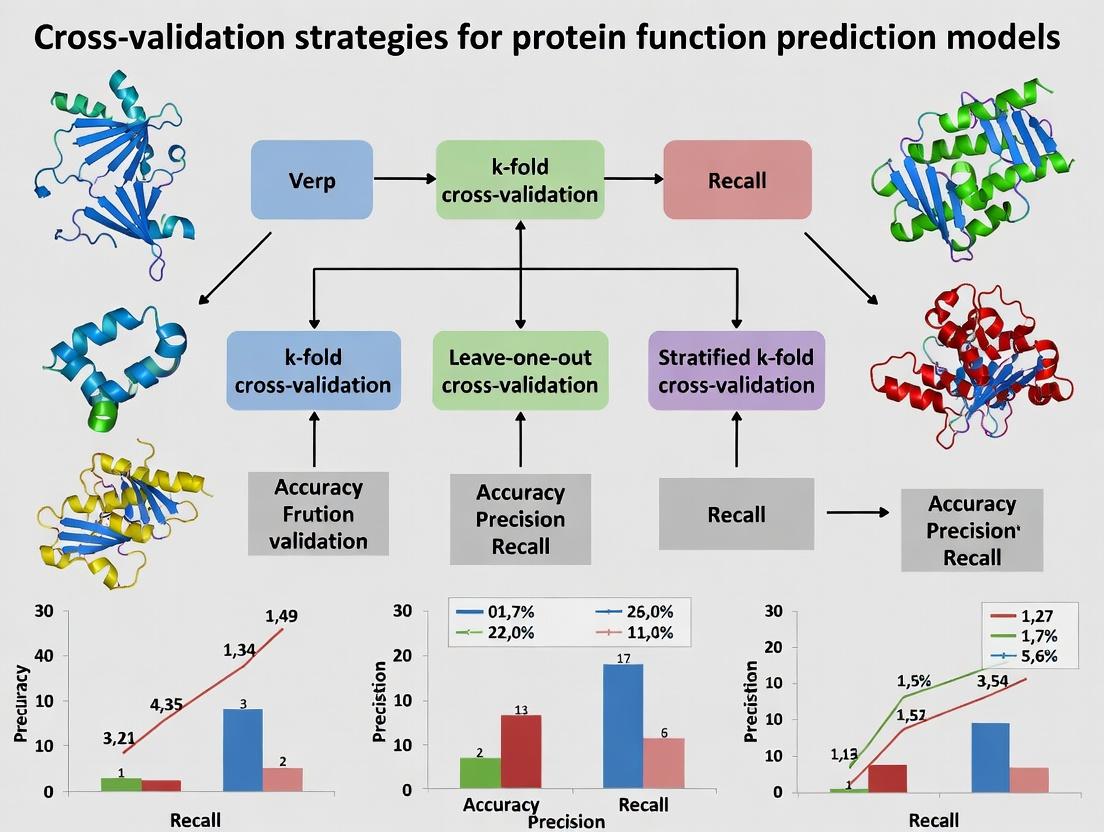
Robust Cross-Validation Strategies for Protein Function Prediction: A Practical Guide for Biomedical AI
This article provides a comprehensive framework for implementing and evaluating cross-validation strategies in protein function prediction models.
Decoding CAPE: A Critical Guide to the Protein Engineering Challenge for Drug Development Researchers
This article provides a comprehensive, critical assessment of the Critical Assessment of Protein Engineering (CAPE) challenge, a pivotal community-wide initiative benchmarking computational tools in protein design.
Contrastive Learning for Protein Representation: A Guide for AI-Driven Drug Discovery
This article provides a comprehensive guide to contrastive learning methods for protein representation, tailored for researchers and drug development professionals.