Protein Language Models vs. BLASTp: A Comparative Guide for Modern Enzyme Annotation
This article provides a comprehensive analysis for researchers and drug development professionals on the evolving landscape of protein function annotation, focusing on the comparative strengths of traditional BLASTp and emerging...
Protein Language Models vs. BLASTp: A Comparative Guide for Modern Enzyme Annotation
Abstract
This article provides a comprehensive analysis for researchers and drug development professionals on the evolving landscape of protein function annotation, focusing on the comparative strengths of traditional BLASTp and emerging protein Language Models (pLMs). We explore the foundational principles of both methods, detail their practical applications in workflows like Enzyme Commission (EC) number prediction, and offer optimization strategies. Drawing on the latest 2025 research, we present a head-to-head validation of their performance, particularly for difficult-to-annotate enzymes. The conclusion synthesizes key takeaways and future directions, advocating for a synergistic approach that leverages the precision of BLASTp with the powerful, homology-independent pattern recognition of pLMs to accelerate biomedical discovery.
From Sequence Alignment to Semantic Understanding: The Foundations of BLASTp and Protein LLMs
In the field of bioinformatics, accurately annotating protein function is a fundamental task that enables researchers to decipher biological processes, understand disease mechanisms, and identify drug targets. For decades, the gold standard tool for this task has been BLASTp (Basic Local Alignment Search Tool for protein sequences), which operates on the fundamental principle that enzymes sharing high sequence similarity likely have similar functions [1]. This homology-based approach has served as the backbone of genome annotation pipelines, allowing for the functional transfer of annotations from well-characterized proteins to novel sequences based on evolutionary relationships. However, the rapid emergence of protein language models (pLMs) like ESM2, ESM1b, and ProtBERT, based on deep learning transformer architectures, presents a powerful new paradigm for function prediction that does not rely exclusively on sequence homology [2] [3]. These models, pre-trained on millions of protein sequences, can learn complex patterns and structural features that extend beyond direct evolutionary relationships. This guide provides an objective comparison of these competing approaches, examining their relative performance through recent experimental data to help researchers and drug development professionals navigate the evolving landscape of protein annotation tools.
Methodological Foundations: How BLASTp and Protein LLMs Work
BLASTp: Sequence Alignment and Homology Transfer
The operational principle of BLASTp is straightforward: it takes a query protein sequence and searches it against a reference database of proteins with known functions. Using a heuristic algorithm, it identifies regions of local similarity between the query and database sequences. The key assumption is that sequence similarity implies evolutionary descent from a common ancestor (homology), which in turn implies functional similarity [1]. The tool then transfers the functional annotation—such as an Enzyme Commission (EC) number—from the best-matching sequence(s) in the database to the query sequence. This method is computationally efficient and biologically intuitive, explaining its enduring popularity. Notably, the National Center for Biotechnology Information (NCBI) has continuously enhanced BLASTp, announcing that by August 2025, the default database will transition to ClusteredNR, which groups redundant sequences to provide faster searches, decreased redundancy, and broader taxonomic coverage in results [4].
Protein Language Models: Learning Sequence Semantics
Protein language models represent a fundamentally different approach. Inspired by successes in natural language processing, pLMs treat protein sequences as "sentences" where amino acids are the "words." Models like ESM2 and ProtBERT are pre-trained on massive datasets of protein sequences (e.g., from UniProtKB) using self-supervised objectives, such as predicting masked amino acids in sequences [3] [5]. Through this process, they learn deep contextual representations of protein sequences, capturing information about biochemical properties, evolutionary constraints, and even structural features without explicit supervision. For downstream tasks like EC number prediction, these learned representations (embeddings) are extracted and used as input features for classifiers, typically fully connected neural networks, which learn to map the embeddings to functional classes [2] [1]. This approach allows pLMs to identify functional signatures even in the absence of close homologs.
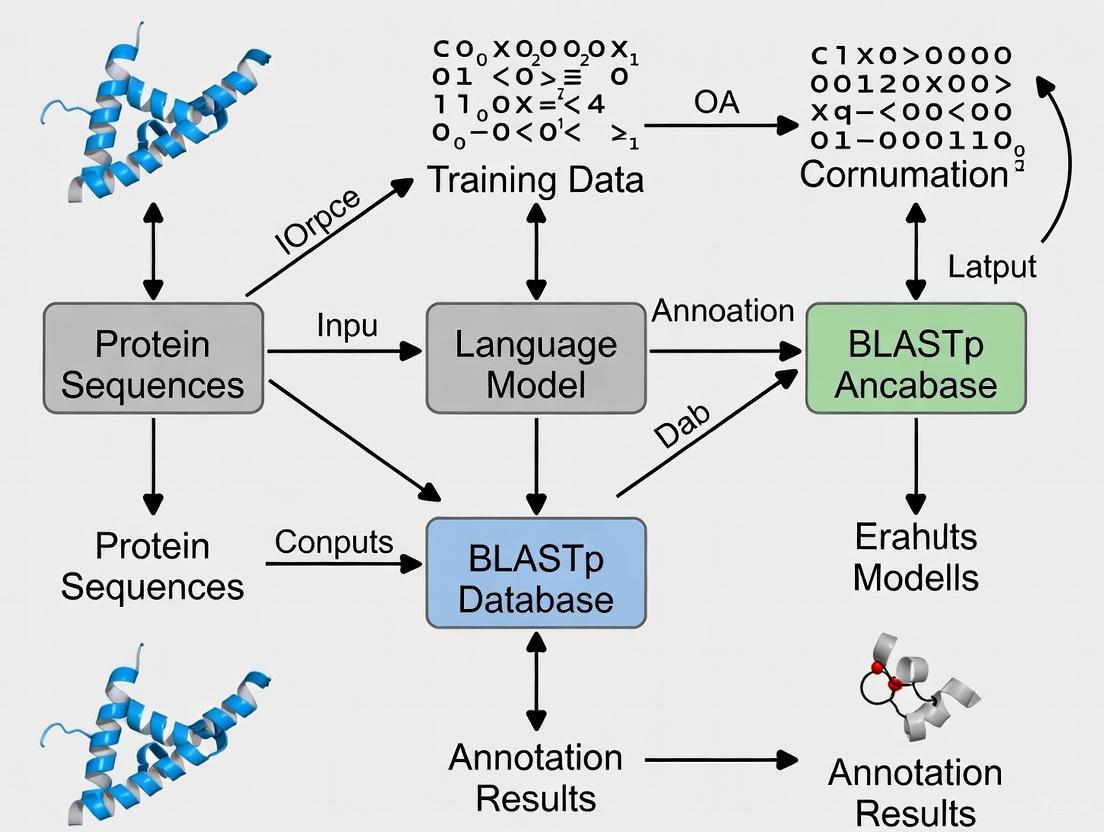
Visualizing the Core Annotation Paradigms
The following diagram illustrates the fundamental differences in how BLASTp and protein Language Models approach the problem of protein function annotation.
Experimental Comparison: Performance Metrics and Limitations
Direct Performance Comparison on EC Number Prediction
A comprehensive 2025 study directly compared the performance of BLASTp against several protein language models (ESM2, ESM1b, and ProtBERT) for predicting Enzyme Commission (EC) numbers, providing robust quantitative data on their relative strengths and weaknesses [2] [1]. The experimental protocol involved training deep learning models using embeddings from each pLM as features, with BLASTp serving as the baseline. The models were evaluated on their ability to correctly assign EC numbers in a multi-label classification setting, incorporating both promiscuous and multi-functional enzymes. The test datasets were constructed from UniProtKB, using only UniRef90 cluster representatives to ensure sequence diversity and avoid overfitting [1].
Table 1: Comparative Performance of BLASTp vs. Protein Language Models for EC Number Prediction
| Method | Overall Accuracy | Strength on High-Identity Sequences (>25%) | Performance on Low-Identity Sequences (<25%) | Ability to Annotate Orphans (No Homologs) |
|---|---|---|---|---|
| BLASTp | Marginally Better | Excellent | Limited | None |
| ESM2 (Best pLM) | Slightly Lower | Good | Good | Yes |
| ESM1b | Lower | Moderate | Moderate | Limited |
| ProtBERT | Lower | Moderate | Moderate | Limited |
The results demonstrated that while BLASTp provided marginally better results overall, the deep learning models provided complementary results, with each method excelling on different subsets of EC numbers [2]. Specifically, the ESM2 model stood out as the best performer among the pLMs, providing more accurate predictions on difficult annotation tasks and for enzymes without homologs in reference databases [1]. Crucially, the study concluded that pLMs still require further improvement to replace BLASTp as the gold standard in mainstream enzyme annotation routines, but they already offer valuable capabilities for specific challenging cases where traditional homology-based methods fail [2].
Performance on Challenging Cases and Low-Similarity Sequences
One of the most significant findings from recent comparative studies is the complementary nature of these approaches, particularly when dealing with sequences that have low similarity to proteins in reference databases. While BLASTp struggles when sequence identity falls below 25%, protein language models maintain reasonable predictive accuracy even for these difficult cases [1]. This capability is particularly valuable for annotating orphan enzymes—those with no recognizable homologs in current databases—which represent a significant challenge in genome annotation projects. The ESM2 model specifically demonstrated robust performance on these difficult annotation tasks, suggesting that pLMs capture functional signals beyond what is accessible through direct sequence comparison alone [2]. This complementary performance profile has led researchers to suggest that hybrid approaches combining both methods may offer the most robust solution for comprehensive genome annotation.
Next-Generation Annotation Systems: Integrating Multiple Approaches
The evolution of bacterial genome annotation systems illustrates the growing trend toward integrating multiple annotation methodologies. BASys2, a next-generation bacterial genome annotation system released in 2025, leverages over 30 bioinformatics tools and 10 different databases to achieve unprecedented annotation depth—generating up to 62 annotation fields per gene/protein while reducing annotation time from 24 hours to as little as 10 seconds [6]. While still relying on BLAST for certain annotation transfers, systems like BASys2 represent a move toward more comprehensive pipelines that can incorporate diverse prediction methods, including emerging deep learning approaches, to provide richer functional insights beyond what any single method can deliver.
Table 2: Key Research Tools for Protein Function Annotation
| Tool/Resource | Type | Primary Function | Application Context |
|---|---|---|---|
| BLASTp | Sequence Alignment Tool | Identifies similar sequences in databases | Primary homology-based annotation |
| ClusteredNR | Protein Database | Non-redundant clustered reference database | Default BLASTp database (from Aug 2025) |
| ESM2 | Protein Language Model | Generates embeddings for function prediction | EC prediction without close homologs |
| ProtBERT | Protein Language Model | Transformer-based sequence representations | Alternative pLM for feature extraction |
| UniProtKB | Protein Knowledgebase | Manually/automatically annotated sequences | Training data source for pLMs |
| FANTASIA | Annotation Tool | Functional annotation based on embedding similarity | Large-scale proteome annotation with pLMs |
The comparative assessment of BLASTp and protein language models reveals a nuanced landscape where neither approach completely dominates the other. BLASTp maintains its position as the gold standard for routine annotation due to its marginally superior overall accuracy, computational efficiency, and deep integration into established bioinformatics workflows [2] [1]. Its upcoming transition to the ClusteredNR database will further enhance its performance by reducing redundancy and providing broader taxonomic coverage [4]. However, protein language models, particularly ESM2, have demonstrated compelling capabilities for addressing challenging annotation scenarios where traditional homology-based methods falter—especially for sequences with low similarity to known proteins and orphan enzymes without database homologs [2] [1]. Rather than viewing these approaches as mutually exclusive, the evidence suggests they offer complementary strengths. Forward-looking researchers and drug development professionals would be well-served by developing workflows that strategically employ both methodologies, leveraging BLASTp for high-confidence homology-based annotations while reserving protein language models for the growing subset of proteins that defy conventional classification through sequence similarity alone. As both technologies continue to evolve—with BLASTp benefiting from database optimizations and pLMs advancing through architectural improvements and training on larger datasets—their synergistic integration promises to push the boundaries of what's possible in protein function prediction.
The emerging paradigm of protein language models (pLMs) represents a fundamental shift in how computational biology approaches the central challenge of linking protein sequence to function. Inspired by breakthroughs in natural language processing, this new framework treats amino acid sequences as sentences in a foreign language, allowing models to learn the complex "grammar" that governs protein structure and function directly from unlabeled sequence data [7]. This approach marks a significant departure from traditional, homology-based methods like BLASTp, which have served as the gold standard for decades by transferring functional annotations from evolutionarily related proteins [8]. Where BLASTp operates on the principle of explicit sequence comparison, protein LLMs such as ESM2, ESM1b, and ProtBERT learn implicit patterns and biochemical constraints from millions of sequences, enabling them to make functional predictions even for proteins without clear homologs in existing databases [2] [9].
This comparative guide examines the performance of these two competing paradigms—the established homology-based approach and the emerging AI-driven framework—within the specific context of enzyme function prediction. By objectively evaluating experimental data on their relative strengths and limitations, we provide researchers, scientists, and drug development professionals with the evidence needed to select appropriate tools for their functional annotation workflows and to understand where the field is heading in the coming years.
Performance Comparison: pLMs vs. Traditional Methods
Quantitative Performance Benchmarks
Direct comparative studies reveal a nuanced performance landscape where traditional and AI-based methods each display distinct advantages depending on the specific prediction context.
Table 1: Performance Comparison of Protein LLMs vs. BLASTp for EC Number Prediction
| Method | Overall Accuracy | Performance on Low-Homology Sequences (<25% identity) | Key Strengths | Limitations |
|---|---|---|---|---|
| BLASTp | Marginally better overall [2] | Significant performance decrease [2] | Excellent for sequences with clear homologs [2] | Cannot annotate orphan sequences without homologs [10] |
| ESM2 (Best-performing pLM) | Slightly lower than BLASTp but complementary [2] | Superior performance on difficult-to-annotate enzymes [2] [10] | Predicts functions for sequences without homologs [2] | Not yet gold standard for mainstream annotation [2] |
| ESM1b | Lower than ESM2 [2] | Good performance on low-homology targets [2] | Useful feature extraction for function prediction [9] | Not state-of-the-art among pLMs [2] |
| ProtBERT | Lower than ESM2 [2] | Moderate performance on low-homology targets [2] | Can be fine-tuned for specific prediction tasks [10] | Underperforms compared to ESM models in benchmarks [2] |
Performance Across Different Prediction Contexts
Beyond enzyme commission number prediction, the relative performance of these methods varies across different functional annotation tasks:
Table 2: Method Performance Across Protein Analysis Tasks
| Task | Best Performing Methods | Key Findings |
|---|---|---|
| Gene Ontology (GO) Term Prediction | BLASTp, MMseqs2, DIAMOND (with optimal parameters) [8] | BLASTp and MMseqs2 consistently exceed other tools under default parameters [8] |
| Protein-Protein Interaction Prediction | SWING (Specialized Interaction Language Model) [11] | Specialized interaction language models outperform generic pLM embeddings for PPI prediction [11] |
| Structure Prediction | AlphaFold-Multimer, DeepSCFold [12] | Integration of sequence-based deep learning with co-evolutionary signals yields highest accuracy [12] |
Experimental Protocols and Methodologies
Standardized Evaluation Framework for EC Number Prediction
To enable fair comparison between traditional and AI-based methods, recent studies have established rigorous experimental protocols. The key methodology for benchmarking EC number prediction involves:
Data Preparation and Processing:
- Source Data: Protein sequences and EC number annotations are extracted from UniProtKB, encompassing both SwissProt (manually annotated) and TrEMBL (automatically annotated) databases [10].
- Sequence Clustering: To avoid homology bias, sequences are clustered using UniRef90, which groups sequences that share at least 90% identity. Only cluster representatives are used in training and testing sets [10].
- Problem Formulation: EC number prediction is treated as a multi-label classification problem, accounting for promiscuous and multi-functional enzymes with multiple EC numbers. The hierarchical nature of EC numbers is preserved, with classifiers challenged to predict the entire hierarchy of labels and their relationships [10].
Model Training and Architecture:
- pLM-Based Models: Protein LLMs (ESM2, ESM1b, ProtBERT) are used as feature extractors, generating embeddings from protein sequences. These embeddings are then fed into fully connected neural networks for EC number classification [2] [10].
- Baseline DL Models: Deep learning models like DeepEC and D-SPACE that rely on one-hot encodings of amino acid sequences are implemented for comparison [10].
- Traditional Methods: BLASTp searches are performed against reference databases with function transfer based on highest-scoring hits [2].
Evaluation Metrics:
- Standard classification metrics including precision, recall, F1-score, and area under the receiver operating characteristic curve [2].
- Stratified evaluation based on sequence similarity thresholds (e.g., <25% identity) to assess performance on low-homology targets [2].
- per-class performance analysis to identify which EC classes are better predicted by each method [2].
The "Protein-as-Second-Language" Framework
An emerging experimental approach reformulates protein function prediction as a zero-shot learning problem:
- Framework Design: Amino acid sequences are treated as sentences in a novel symbolic language. The framework adaptively constructs sequence-question-answer triples that reveal functional cues without task-specific training [13].
- Data Curation: A bilingual corpus of 79,926 protein-QA instances spanning attribute prediction, descriptive understanding, and extended reasoning is created to support this approach [13].
- Evaluation: The method is tested in a zero-shot setting across diverse LLMs, achieving improvements of up to 17.2% ROUGE-L (average +7%) and even surpassing fine-tuned protein-specific language models in some cases [13].
Workflow Visualization: From Sequence to Function
Traditional vs. pLM-Based Annotation Pathways
The fundamental differences between traditional homology-based approaches and the new pLM paradigm can be visualized through their distinct workflows:
pLM Architecture and Training Methodology
Protein language models learn the "grammar" of proteins through self-supervised training on massive sequence datasets:
Essential Research Reagents and Computational Tools
The Scientist's Toolkit for Protein Function Annotation
Table 3: Key Research Reagents and Computational Tools for Protein Function Prediction
| Tool/Resource | Type | Primary Function | Application Context |
|---|---|---|---|
| ESM2 [2] [10] | Protein Language Model | Feature extraction from protein sequences | State-of-the-art for EC number prediction; best-performing among pLMs |
| ProtBERT [2] [10] | Protein Language Model | Feature extraction and fine-tuning for specific tasks | EC number prediction; can be fine-tuned for specific prediction tasks |
| BLASTp [2] [8] | Sequence Alignment Tool | Homology-based function transfer | Gold standard for sequences with clear homologs; widely used in annotation pipelines |
| DIAMOND [8] | Sequence Alignment Tool | Fast homology search | BLASTp alternative optimized for speed with slightly lower sensitivity |
| MMseqs2 [8] | Sequence Alignment Tool | Fast, sensitive sequence search | Performance comparable to BLASTp with correct parameter settings |
| SWING [11] | Specialized Interaction Model | Protein-protein interaction prediction | Outperforms generic pLMs for interaction-specific tasks |
| UniProtKB [10] | Protein Database | Source of annotated sequences | Primary data source for training and benchmarking |
| UniRef90 [10] | Clustered Protein Database | Sequence similarity-based clustering | Reduces homology bias in training datasets |
The experimental evidence clearly demonstrates that protein language models and traditional homology-based methods represent complementary rather than strictly competitive approaches to protein function annotation. While BLASTp maintains a marginal overall advantage for routine annotation of proteins with clear evolutionary relatives [2], protein LLMs excel in the critical task of predicting functions for difficult-to-annotate enzymes, particularly when sequence identity falls below 25% [2] [10]. This performance profile suggests an integrated future for protein function prediction, where LLMs handle the challenging cases that evade traditional homology-based methods while BLASTp continues to provide reliable annotations for sequences with clear homologs.
The most effective annotation pipelines will likely leverage both approaches, combining the evolutionary signals captured by traditional methods with the learned biochemical constraints embedded in protein LLMs. As these models continue to evolve—with emerging frameworks treating "protein as a second language" for LLMs [13] and specialized interaction language models like SWING [11] addressing specific prediction tasks—the gap between sequence and function will continue to narrow, accelerating discovery in basic research and drug development alike.
The field of protein function prediction has undergone a fundamental transformation, moving from traditional similarity-based methods toward deep learning approaches that capture complex biological patterns. For decades, BLASTp has served as the gold standard for protein annotation, operating on the principle that proteins with similar sequences share similar functions [3]. While effective for detecting clear homologs, this approach struggles with remote homology and fails to leverage the full contextual information embedded in protein sequences. The advent of protein Language Models (pLMs), built on Transformer architectures and self-attention mechanisms, represents a paradigm shift. These models, pre-trained on millions of protein sequences, learn the underlying "language of life," capturing intricate biochemical and structural properties that enable more accurate and generalizable function prediction, even for proteins with low sequence similarity to known proteins [14] [3].
This guide provides an objective comparison of these competing methodologies, focusing on their core architectures, performance benchmarks, and practical applications in bioinformatics and drug development. We present experimental data from recent, comprehensive studies to help researchers select the appropriate tool for their protein annotation needs.
Core Architectural Differences: BLASTp vs. Transformer-based pLMs
The fundamental difference between these approaches lies in how they process and interpret protein sequence information.
BLASTp: A Sequence Alignment Workhorse
BLASTp (Basic Local Alignment Search Tool for proteins) employs a local alignment strategy to identify regions of similarity between a query sequence and a database of known proteins. Its methodology is based on heuristics to rapidly find sequence matches, after which it estimates the statistical significance of these matches (E-values) [1]. The underlying assumption is that function can be transferred from a well-annotated protein to a query protein based on significant sequence similarity. While recent database improvements like ClusteredNR reduce redundancy and improve search speed, the core algorithm remains based on pairwise sequence comparison [4].
Transformer-based pLMs: Learning the Language of Proteins
In contrast, pLMs like ESM-2 and ProtT5 are based on the Transformer architecture. At their core is the self-attention mechanism, which allows the model to weigh the importance of all amino acids in a sequence when representing any single residue. This enables the model to capture long-range dependencies and complex interactions that are invisible to local alignment [14] [15].
These models are first pre-trained on massive datasets (e.g., UniRef) using self-supervised objectives like Masked Language Modeling (MLM), where the model learns to predict randomly masked amino acids based on their context. The resulting model contains rich, contextual representations of protein sequences, which can then be fine-tuned for specific downstream tasks such as protein-protein interaction prediction, enzyme classification, or crystallization propensity prediction [16] [17].
Table: Core Architectural Comparison between BLASTp and Transformer-based pLMs
| Feature | BLASTp | Transformer-based pLMs |
|---|---|---|
| Underlying Principle | Local sequence alignment & homology transfer | Contextual sequence representation via self-supervised learning |
| Core Mechanism | Heuristic search for similar sequence segments | Self-attention mechanism capturing residue-residue dependencies |
| Training Data | Reference protein databases (e.g., nr, ClusteredNR) | Large, unlabeled sequence corpora (e.g., UniRef) |
| Primary Output | Sequence matches with statistical significance (E-values) | Contextual embeddings for each residue and/or the entire protein |
| Key Strength | Excellent for finding clear homologs; intuitive interpretation | Superior for detecting remote homology & capturing functional signatures |
Performance Benchmarking: Experimental Data and Comparisons
Enzyme Commission (EC) Number Prediction
A comprehensive 2025 study directly compared the performance of pLMs and BLASTp for predicting enzyme functions, defined by their EC numbers [1]. The research evaluated several pLMs, including ESM2, ESM1b, and ProtBERT, against BLASTp on a large test set. The results revealed that while BLASTp provided marginally better overall results, the pLM-based models offered complementary performance, with each method excelling on different subsets of EC numbers [1].
Crucially, the study found that pLMs significantly outperformed BLASTp for enzymes where the identity between the query sequence and the reference database fell below 25%. This highlights the particular value of pLMs for annotating proteins with few or distant homologs. Among the pLMs, ESM2 stood out as the most effective, providing more accurate predictions for difficult annotation tasks [1].
Table: Performance Comparison in EC Number Prediction [1]
| Method | Overall Performance | Performance on Low-Identity Sequences (<25% Identity) | Key Strengths |
|---|---|---|---|
| BLASTp | Marginal overall advantage | Lower performance | Best for proteins with high-identity homologs |
| ESM2 (pLM) | Competitive, slightly lower overall | Significantly better performance | Superior for remote homology, difficult annotations |
| ESM1b (pLM) | Lower than ESM2 and BLASTp | Moderate | - |
| ProtBERT (pLM) | Lower than ESM2 and BLASTp | Moderate | - |
Protein-Protein Interaction (PPI) Prediction
The application of pLMs has expanded beyond single-protein annotation to predicting interactions between proteins. A 2025 study introduced PLM-interact, a model based on a fine-tuned ESM-2 architecture, specifically designed for PPI prediction [16]. The model was trained on human PPI data and tested on data from five other species (mouse, fly, worm, yeast, and E. coli) in a rigorous cross-species benchmark.
PLM-interact achieved state-of-the-art performance, outperforming six other PPI prediction methods (TUnA, TT3D, Topsy-Turvy, D-SCRIPT, PIPR, and DeepPPI) in most tested scenarios [16]. The performance improvement was particularly notable for the more challenging yeast and E. coli datasets, where PLM-interact achieved a 10% and 7% improvement in AUPR (Area Under the Precision-Recall Curve), respectively, over the next best method (TUnA) [16]. This demonstrates the power of transformer models to generalize learned interaction patterns across evolutionary distances.
Protein Crystallization Propensity Prediction
A 2025 benchmarking study evaluated various pLMs for predicting a protein's propensity to form crystals, a critical step in structural biology [17]. The research compared classifiers built on embedding representations from models including ESM2, Ankh, ProtT5-XL, ProstT5, xTrimoPGLM, and SaProt.
The study found that LightGBM classifiers utilizing embeddings from ESM2 models (with 30 and 36 transformer layers) outperformed all other sequence-based methods, including DeepCrystal, ATTCrys, and CLPred [17]. These ESM2-based predictors achieved performance gains of 3-5% in key metrics like AUPR, AUC, and F1-score on independent test sets, demonstrating the practical utility of pLM embeddings for challenging biophysical property prediction [17].
Experimental Protocols and Methodologies
Standard Protocol for pLM-based Function Prediction
A typical experimental pipeline for using pLMs in protein function prediction involves several key stages, as detailed in comparative studies [1] [17]:
- Sequence Input and Preprocessing: Protein amino acid sequences are obtained in FASTA format. Sequences are often filtered by length and quality.
- Embedding Generation: Sequence(s) are passed through a pre-trained pLM (e.g., ESM2) to generate numerical representations (embeddings). These can be per-residue embeddings or a single, pooled representation for the entire protein.
- Downstream Model Training: The generated embeddings are used as input features for a machine learning classifier (e.g., a fully connected neural network, LightGBM, or XGBoost) trained to predict specific functional labels (e.g., EC numbers, GO terms).
- Performance Evaluation: The model is evaluated on held-out test sets using standard metrics such as AUROC, AUPR, and F1-score. Cross-validation is often employed to ensure robustness.
Protocol for Cross-Species PPI Prediction with PLM-interact
The PLM-interact model demonstrated a specialized architecture and training regimen for interaction prediction [16]:
- Architecture Modification: Starts with a pre-trained ESM-2 model and extends it to jointly encode protein pairs. This is analogous to the "next-sentence prediction" task in natural language processing.
- Balanced Training: The model is fine-tuned using a combined loss function that balances the Masked Language Modeling (MLM) objective with a Next Sentence Prediction (classification) objective. A specific 1:10 ratio between classification and mask loss was found to be optimal.
- Training Data: The model is trained on large datasets of known interacting and non-interacting protein pairs, such as 421,792 human protein pairs.
- Cross-Species Validation: The model's generalizability is tested by evaluating its performance on protein interaction data from species not seen during training (e.g., training on human, testing on mouse, fly, worm, yeast, E. coli).
The following tools and databases are critical for researchers working in the field of protein annotation and function prediction.
Table: Essential Research Tools for Protein Annotation
| Tool Name | Type | Primary Function | Relevance |
|---|---|---|---|
| ESM-2 [16] [15] | Protein Language Model | Generates contextual embeddings from protein sequences | State-of-the-art pLM for various downstream tasks |
| TRILL [17] | Computational Platform | Democratizes access to multiple pLMs for property prediction | Allows easy benchmarking of different pLMs without deep coding expertise |
| ClusteredNR [4] | Protein Database | Non-redundant clustered protein database for BLAST | Reduces redundancy and speeds up BLAST searches |
| BASys2 [6] | Annotation Pipeline | Rapid, comprehensive bacterial genome annotation | Integrates >30 tools for functional and structural annotation |
| PLM-interact [16] | Specialized pLM | Predicts protein-protein interactions from sequence | Demonstrates extension of pLMs to complex relational tasks |
| CARD [18] | Specialized Database | Curated database of antimicrobial resistance genes | Essential for AMR annotation; used in minimal model benchmarking |
The experimental evidence clearly demonstrates that transformer-based pLMs and traditional BLASTp offer complementary strengths. BLASTp remains a robust, efficient tool for annotating proteins with clear, high-identity homologs. Its interpretability and speed make it a valuable first pass in many annotation pipelines.
However, pLMs have established a dominant advantage in scenarios involving remote homology, complex functional patterns, and predictions where pure sequence similarity is low. Their ability to learn the intricate "grammar" of protein sequences from unlabeled data allows them to uncover functional insights that evade alignment-based methods. As the field progresses, the most powerful annotation pipelines will likely strategically combine both approaches, leveraging the respective strengths of each method to achieve maximum accuracy and coverage [1]. Future developments will focus on increasingly specialized pLMs, improved interpretability of attention mechanisms [15], and integration of structural information for even more powerful protein function prediction.
The accurate prediction of enzyme function, classified by Enzyme Commission (EC) numbers, is a critical task in bioinformatics with profound implications for understanding cellular metabolism, drug discovery, and genome annotation. For decades, similarity-based search tools like BLASTp have served as the gold standard for this function. However, the recent emergence of protein Large Language Models (pLMs)—including ESM2, ESM1b, and ProtBERT—offers a powerful new paradigm for extracting functional insights directly from sequence data. This guide provides a objective, data-driven comparison of these three prominent pLMs, benchmarking their performance against each other and traditional BLASTp-based annotation to inform researchers and drug development professionals about their respective capabilities and optimal use cases.
The following tables summarize the key performance characteristics and experimental results for ESM2, ESM1b, and ProtBERT in the context of EC number prediction.
Table 1: Model Architectures and Key Performance Insights
| Model | Key Architecture/Pre-training | Overall Performance vs. BLASTp | Key Strength | Notable Limitation |
|---|---|---|---|---|
| ESM2 | Transformer; pre-trained on UniProtKB [1] | Best among pLMs; slightly behind BLASTp overall [1] | Most accurate for difficult annotations & enzymes without homologs [1] | - |
| ESM1b | Transformer; pre-trained on UniProtKB [1] [3] | Surpassed by ESM2 [1] | Widely applied for improving prediction accuracy [3] | Outperformed by newer ESM2 variants [1] |
| ProtBERT | Transformer; pre-trained on UniProtKB & BFD [1] | Surpassed by ESM2 [1] | Often fine-tuned for EC prediction [1] | - |
| BLASTp | Local sequence alignment & homology search [1] | Marginally better overall results than pLMs [1] | Excels for many EC numbers with clear homologs [1] | Cannot annotate proteins without homologous sequences [1] |
Table 2: Comparative Experimental Data on EC Number Prediction
| Metric / Characteristic | ESM2 | ESM1b | ProtBERT | BLASTp |
|---|---|---|---|---|
| Performance with Low Identity (<25%) | Good predictions [1] | Information missing | Information missing | Performance decreases [1] |
| Complementarity with BLASTp | Yes - predicts EC numbers that BLASTp misses [1] | Implied by category | Implied by category | Yes - predicts EC numbers that pLMs miss [1] |
| Input Representation | Embeddings (Feature Extraction) [1] | Embeddings (Feature Extraction) [1] | Embeddings (often Fine-tuning) [1] | Amino Acid Sequence (Direct Search) |
| Optimal Combined Use | More effective when used together with BLASTp [1] | More effective when used together with BLASTp [1] | More effective when used together with BLASTp [1] | More effective when used together with pLMs [1] |
Experimental Protocols and Methodology
A robust experimental framework is essential for a fair comparison of protein function prediction tools. The following workflow and detailed methodology outline the standard approach for benchmarking EC number prediction performance.
Experimental Workflow for EC Number Prediction
Detailed Methodological Components
Data Sourcing and Processing
The standard protocol utilizes protein data from UniProtKB (SwissProt and TrEMBL) downloaded in XML format. To ensure data quality and reduce redundancy, only UniRef90 cluster representatives are retained. These representatives are selected based on entry quality, annotation score, organism relevance, and sequence length, creating a non-redundant dataset ideal for model training and evaluation [1].
Problem Formulation
EC number prediction is formally defined as a global hierarchical multi-label classification problem. Each protein sequence is associated with a binary vector indicating all relevant EC numbers across all hierarchical levels (from the first digit to the fourth). This approach accounts for promiscuous and multi-functional enzymes, requiring a single classifier to predict the entire hierarchy of labels and their complex relationships [1].
Feature Extraction and Model Training
- pLM Embeddings: For ESM2, ESM1b, and ProtBERT, the standard approach is feature extraction. The embeddings (numeric representations) from a specific layer of the pre-trained model are used as input features for a downstream classifier. A commonly used and effective compression strategy is mean pooling, which averages embeddings across all amino acid positions in a sequence [5]. These embeddings are then processed by a fully connected neural network to predict the EC number vector [1].
- Baseline Models: For comparison, deep learning models like DeepEC and D-SPACE, which use one-hot encodings of amino acid sequences as input, are implemented. This setup allows for a direct comparison between traditional input representations and those learned by pLMs [1].
- BLASTp Protocol: The BLASTp baseline involves performing a standard similarity search against a curated database of enzymes with known EC numbers. The function is transferred from the best hit(s) based on sequence similarity thresholds [1].
The Scientist's Toolkit: Key Research Reagents and Solutions
Table 3: Essential Resources for Protein Function Prediction Research
| Resource / Tool | Type | Primary Function in Research |
|---|---|---|
| UniProtKB Database | Data Repository | Provides the foundational, curated protein sequences and their functional annotations (including EC numbers) for model training and testing [1]. |
| ESM2 / ESM1b / ProtBERT | Protein Language Model | Serves as the core feature extractor, converting raw amino acid sequences into semantically rich, numerical embeddings (vector representations) for downstream prediction tasks [1]. |
| BLASTp | Bioinformatics Tool | Functions as the standard baseline for performance comparison based on sequence homology and homology-based function transfer [1]. |
| Fully Connected Neural Network | Deep Learning Model | Acts as the final classifier that takes pLM embeddings as input and performs the multi-label classification to assign EC numbers [1]. |
| ClusteredNR Database | Protein Sequence Database | An NCBI database of clustered protein sequences that reduces redundancy. It is becoming the new default for BLASTp, enabling faster searches with broader taxonomic coverage [4]. |
Critical Analysis and Research Implications
The experimental data leads to several critical conclusions for researchers. First, while BLASTp maintains a marginal overall advantage, the performance of pLMs and BLASTp is complementary [1]. Each method excels at predicting different subsets of EC numbers, suggesting that a combined approach is more powerful than either method in isolation.
Second, among the pLMs, ESM2 consistently emerges as the top performer, providing the most accurate predictions for challenging annotation tasks, especially for enzymes with no close homologs or when sequence identity to known proteins falls below 25% [1]. This makes ESM2 particularly valuable for exploring the "microbial dark matter" in metagenomic studies.
Finally, a crucial consideration for practical application is the balance between model size and performance. While larger pLMs exist, recent evidence suggests that medium-sized models (e.g., ESM2 650M) often achieve performance comparable to their much larger counterparts (e.g., ESM2 15B) in many transfer learning scenarios, offering a superior balance of computational efficiency and predictive power [5].
The comparative analysis of ESM2, ESM1b, and ProtBERT reveals a nuanced landscape in protein function prediction. ESM2 currently holds a slight edge among pLMs for EC number prediction, particularly for difficult cases with low sequence homology. However, the longstanding BLASTp tool remains a robust and marginally superior performer overall for routine annotations. The most effective strategy for researchers is not to choose one over the other, but to leverage their complementary strengths. Integrating pLM-based predictions with traditional homology-based methods like BLASTp provides the most comprehensive and accurate path forward for annotating the functional universe of enzymes.
In the era of high-throughput sequencing, functional genomics faces a critical bottleneck: the immense gap between the rapid accumulation of protein sequences and their functional characterization. As of February 2024, the UniProt database contains over 240 million protein sequences, yet less than 0.3% have experimentally validated functional annotations [9]. This staggering discrepancy represents what researchers term the "dark proteome" – a vast landscape of uncharacterized proteins that may hold keys to understanding biological processes, disease mechanisms, and therapeutic targets [19]. For researchers and drug development professionals, this annotation gap presents both a challenge and an imperative: without accurate functional annotations, genomic data remains largely uninterpretable, potentially obscuring valuable insights for drug discovery and basic biological research.
Traditional annotation methods, primarily relying on sequence similarity through tools like BLASTp, have fundamental limitations when dealing with proteins that lack clear homologs in databases [2] [19]. Approximately 30% of proteins in model organisms like Caenorhabditis elegans remain unannotated, with this figure rising to 50% for non-model organisms [19]. The rapid expansion of genomic data from large-scale initiatives such as the Earth BioGenome Project further exacerbates this problem, generating unprecedented volumes of genomic information that demand reliable annotation frameworks extending beyond conventional approaches [19].
This article examines the emerging landscape of annotation tools, with particular focus on the performance comparison between traditional homology-based methods and novel approaches leveraging protein language models (pLMs). We provide experimental data, methodological details, and practical resources to guide researchers in selecting appropriate tools for their functional genomics workflows.
Performance Comparison: pLMs Versus Traditional Methods
Quantitative Performance Metrics
Table 1: Comparative performance of annotation methods for Enzyme Commission (EC) number prediction
| Method | Overall Accuracy | Performance on Sequences <25% Identity | Key Strengths | Limitations |
|---|---|---|---|---|
| BLASTp | Marginally higher overall [2] | Significant performance decrease [2] | Excellent for sequences with clear homologs [2] | Limited for divergent sequences, orphans [2] [19] |
| ESM2 | High (best among pLMs) [2] | Maintains better accuracy on difficult annotations [2] | Predicts function without homologs; handles remote homology [2] | Still requires improvement to surpass BLASTp in routine annotation [2] |
| ProtT5 | High [20] [19] | Good performance on uncharacterized sequences [20] | Balanced performance; used in FANTASIA pipeline [19] | Computational resource requirements [19] |
| One-hot encoding + DL | Lower than pLMs [2] | Poor performance on sequences without homologs [2] | Simple implementation | Limited generalizability; inferior to modern pLMs [2] |
Table 2: Large-scale proteome annotation performance across animal taxa
| Method | Annotation Coverage | Informativeness of Terms | Novel Function Discovery | Computational Efficiency |
|---|---|---|---|---|
| Homology-based (Traditional) | ~50-60% of genes in non-model organisms [19] | Broad but shallow annotations | Limited to known homologs | Fast but incomplete [19] |
| FANTASIA (ProtT5) | Up to ~50% additional proteins annotated [19] | More precise and informative GO terms [19] | Reveals phylum-specific functions [19] | Moderate; scalable to full proteomes [19] |
| BASys2 | Comprehensive (62 annotation fields) [6] | Integrates multiple data types | Focused on metabolite annotation | Extremely fast (0.5 min average) [6] |
Key Insights from Comparative Studies
The comparative assessment reveals that BLASTp still provides marginally better results overall for routine annotation tasks, particularly when clear homologs exist in databases [2]. However, protein language models demonstrate complementary strengths, excelling in predicting certain EC numbers that challenge homology-based methods and maintaining performance on sequences with identity below 25% [2]. This suggests a synergistic relationship rather than outright replacement.
For large-scale proteome annotation, pLM-based approaches demonstrate remarkable advantages. The FANTASIA pipeline, leveraging ProtT5 embeddings, annotates up to 50% of proteins that remain uncharacterized by traditional homology-based methods [19]. This expanded coverage proves particularly valuable for non-model organisms, where homology-based tools fail to annotate nearly half of the genes, especially in less-studied phyla [19].
The ESM2 model stands out as the best performer among pLMs for EC number prediction, providing more accurate predictions on difficult annotation tasks and for enzymes without homologs [2]. Its architecture, trained on millions of diverse protein sequences, captures evolutionary patterns and structural constraints that enable functional inference beyond sequence similarity.
Experimental Protocols and Methodologies
EC Number Prediction Benchmarking
Experimental Objective: To compare the performance of protein language models (ESM2, ESM1b, ProtBERT) with BLASTp and one-hot encoding-based deep learning models for predicting Enzyme Commission numbers [2].
Data Preparation:
- Curated datasets of enzyme sequences with known EC numbers from public databases
- Partitioned sequences into training, validation, and test sets with careful attention to avoiding data leakage
- Generated sequence identity clusters to assess performance across different evolutionary distances
Model Configurations:
- ESM2: Used the 650M parameter model pre-trained on UniRef50, followed by a fully connected neural network classifier
- ESM1b: Implemented the 650M parameter model with similar architecture to ESM2
- ProtBERT: Utilized the BERT-base architecture pre-trained on BFD and UniRef50 datasets
- BLASTp: Conducted search against reference database with E-value threshold of 0.001
- One-hot encoding: Converted sequences to one-hot vectors followed by CNN architecture
Evaluation Metrics:
- Accuracy per EC number level and overall
- Precision-recall curves for different sequence identity thresholds
- Statistical significance testing between method performances [2]
Large-scale Proteome Annotation with FANTASIA
Experimental Objective: To assess the capability of protein language models for annotating complete proteomes across diverse animal taxa [19].
Pipeline Implementation:
- Input: Proteome files in FASTA format from 970 animal species (~23 million genes)
- Preprocessing: Filtered by length and sequence similarity to remove identical sequences
- Embedding Generation: Computed protein embeddings using ProtT5 model
- Similarity Search: Employed embedding similarity against GOA database
- Function Transfer: Predicted GO terms using closest hits or distance-based filtering
- Output: Functional predictions across three GO categories (biological process, molecular function, cellular component)
Validation Approach:
- Compared annotations with homology-based methods for model organisms
- Assessed coverage (proportion of annotated proteins) and informativeness (detail of terms)
- Conducted phylogenetic enrichment analysis to identify lineage-specific functions [19]
FANTASIA Pipeline: From proteome input to functional annotation
Visualization of Key Concepts and Relationships
The Annotation Gap and Tool Evolution
The Genomic Annotation Challenge: From data generation to interpretation
Complementary Strengths of BLASTp and pLMs
Annotation Strategies: Complementary approaches for comprehensive coverage
Table 3: Key resources for functional genomics annotation
| Resource | Type | Primary Function | Application Context |
|---|---|---|---|
| FANTASIA | Annotation pipeline | Large-scale functional annotation using pLM embeddings [19] | Proteome-wide annotation, non-model organisms [19] |
| BASys2 | Bacterial annotation system | Rapid, comprehensive genome annotation with metabolic focus [6] | Bacterial genomics, metabolite annotation [6] |
| ESM2 | Protein language model | Protein sequence representation for downstream tasks [2] | EC number prediction, remote homology detection [2] |
| ProtT5 | Protein language model | Protein sequence embedding generation [20] [19] | Function prediction, embedding similarity searches [19] |
| SegmentNT | DNA foundation model | Nucleotide-resolution genome annotation [21] | Gene element prediction, regulatory element detection [21] |
| PLSDB | Plasmid database | Curated plasmid sequence resource [22] | Plasmid annotation, horizontal gene transfer studies [22] |
The field of functional genomics stands at a transitional point where traditional homology-based methods and emerging AI-driven approaches are converging toward a hybrid future. Current evidence suggests that protein language models still require refinement to become the gold standard over BLASTp in mainstream annotation routines [2]. However, their superior performance on difficult-to-annotate proteins and capacity to illuminate the "dark proteome" make them indispensable for comprehensive genomic interpretation [19].
For research and drug development professionals, practical implementation should consider a combined approach: using BLASTp for sequences with clear homologs while deploying pLM-based tools for orphan genes, rapidly evolving sequences, and non-model organisms. As these tools evolve, they promise to gradually close the annotation gap, transforming our ability to extract biological meaning from genomic sequence and accelerating discoveries in basic biology and therapeutic development.
The integration of pLMs into annotation pipelines like FANTASIA and BASys2 demonstrates the practical viability of these approaches at scale. With the continued expansion of genomic sequencing initiatives, such advanced annotation tools will become increasingly critical for translating genetic information into biological understanding and clinical applications.
Putting Tools to Work: Methodologies and Real-World Applications in Drug Discovery and Annotation
For decades, BLASTp (Basic Local Alignment Search Tool for protein sequences) has served as the cornerstone of bioinformatics workflows, enabling researchers to compare protein sequences against databases to infer functional and evolutionary relationships [23]. Its fundamental principle rests on identifying regions of local similarity between sequences, operating on the paradigm that sequence similarity often implies functional homology [1]. However, the emerging landscape of artificial intelligence has introduced a powerful new paradigm: protein language models (LLMs) like ESM2, ESM1b, and ProtBERT, which learn complex patterns from millions of protein sequences to predict function [1] [3]. This guide presents a comprehensive overview of the BLASTp workflow while contextualizing its performance and applications relative to these modern computational approaches.
Recent comparative studies reveal a nuanced relationship between traditional alignment tools and AI-based methods. Although BLASTp maintains marginal overall superiority in routine enzyme commission (EC) number annotation, protein LLMs demonstrate complementary strengths, particularly for difficult-to-annotate enzymes and sequences with low homology to known proteins [1] [2]. This evolving dynamic underscores the importance of understanding BLASTp's methodology, optimal implementation, and position within a modern bioinformatics toolkit that increasingly integrates multiple computational strategies.
The BLASTp Workflow: A Step-by-Step Protocol
The standard BLASTp workflow transforms a query protein sequence into functional hypotheses through a series of structured computational steps. The following diagram maps this logical progression from input to biological interpretation:
Input and Database Selection
The process initiates with the query protein sequence in FASTA format. Critical to success is selecting an appropriate protein database for comparison:
- nr (non-redundant): A comprehensive protein database incorporating multiple sources, though soon to be replaced as the BLASTp default [4].
- ClusteredNR: The forthcoming default database (effective August 2025), offering decreased redundancy and broader taxonomic coverage by clustering similar sequences and selecting representative members [4].
- SwissProt: A manually annotated, high-quality database with reduced false positives due to expert curation [1].
Execution and Algorithmic Core
BLASTp employs a heuristic search algorithm that balances sensitivity with computational speed. The process identifies High-scoring Segment Pairs (HSPs) through three core stages:
- Seed Matching: The algorithm identifies short, exact matches ("words") between the query and database sequences.
- Extension: Promising seeds are extended in both directions to form longer alignments, calculating alignment scores.
- Evaluation: Extended alignments are evaluated for statistical significance [23].
Recent BLAST+ 2.17.0 releases have enhanced performance, including faster blastp searches with the -task blastp-fast option and support for compressed FASTA files [24].
Results Interpretation and Statistical Analysis
Proper interpretation requires understanding key metrics that evaluate alignment quality and biological relevance:
- E-value (Expectation Value): The number of alignments expected by chance. Lower E-values (closer to zero) indicate greater statistical significance.
- Bit Score: A normalized score representing alignment quality, independent of database size. Higher scores indicate better matches.
- Percent Identity: The percentage of identical residues in the alignment, providing a straightforward measure of sequence conservation.
- Query Coverage: The percentage of the query sequence included in the alignment, indicating the extent of similarity.
BLASTp Versus Protein Language Models: Experimental Comparisons
Methodology for Comparative Performance Assessment
Recent studies have established rigorous experimental frameworks to evaluate BLASTp against protein LLMs for function prediction, specifically for annotating Enzyme Commission (EC) numbers [1]. The standard protocol involves:
Dataset Preparation:
- Source protein sequences and corresponding EC numbers from UniProtKB (SwissProt and TrEMBL) [1].
- Apply clustering (e.g., UniRef90) to reduce sequence redundancy, ensuring non-homologous training and test sets [1].
- Formulate EC number prediction as a multi-label classification problem accommodating promiscuous and multi-functional enzymes [1].
Model Implementation and Comparison:
- BLASTp: Perform sequence similarity searches against reference databases, transferring EC numbers from top hits based on highest similarity [1].
- Protein LLMs: Extract sequence representations (embeddings) from pre-trained models (ESM2, ESM1b, ProtBERT) and train fully connected neural networks for EC number prediction [1].
- Evaluation Metric: Use precision-recall analysis and accuracy measurements across different EC number hierarchy levels and sequence identity thresholds [1].
Comparative Performance Data
The table below summarizes quantitative findings from a 2025 comparative assessment:
Table 1: Performance Comparison of BLASTp versus Protein Language Models for EC Number Prediction
| Method | Overall Accuracy | Strength Scenarios | Weakness Scenarios | Computational Demand |
|---|---|---|---|---|
| BLASTp | Marginally Higher | High-identity sequences (>25-30% identity) [1] | Sequences with no homologs in databases [1] | Lower (heuristic algorithm) |
| Protein LLMs (ESM2) | Slightly Lower but Complementary | Low-identity sequences (<25% identity), difficult-to-annotate enzymes [1] [2] | Routine annotation of high-similarity sequences [1] | Higher (neural network inference) |
| Hybrid Approach | Highest Reported | Combines strengths of both methods [1] [2] | Implementation complexity | Highest (multiple systems) |
The experimental data reveals that ESM2 emerged as the top-performing protein LLM, providing more accurate predictions for challenging annotation tasks, particularly when sequence identity to known proteins falls below 25% [1] [2]. This suggests that protein LLMs learn functional patterns that extend beyond simple sequence homology.
Essential Research Reagents and Computational Tools
Successful implementation of sequence comparison and analysis requires specific computational tools and resources. The following table catalogs key components for BLASTp and protein LLM workflows:
Table 2: Research Reagent Solutions for Protein Sequence Analysis
| Tool/Resource | Type | Primary Function | Access |
|---|---|---|---|
| BLAST+ Suite | Software Package | Command-line execution of BLASTp searches and database formatting [24] | Free from NCBI |
| ClusteredNR Database | Protein Database | Non-redundant clustered database for reduced redundancy in results [4] | Free from NCBI |
| UniProtKB | Protein Database | Source of expertly curated (SwissProt) and automated (TrEMBL) sequences for training and validation [1] | Free from EMBL-EBI |
| ESM2 Model | Protein Language Model | State-of-the-art protein LLM for generating sequence embeddings and function prediction [1] | Free from Meta AI |
| PyMOL | Visualization Software | Molecular visualization system for structural analysis of query proteins and hits [25] | Commercial (academic pricing) |
Integrated Workflow for Modern Protein Annotation
The complementary strengths of BLASTp and protein language models suggest an integrated approach for comprehensive protein function prediction. The following workflow leverages both methodologies for robust annotation:
This integrated pathway begins with conventional BLASTp analysis, which remains highly effective for sequences with clear homology in reference databases. For sequences lacking significant database matches, or when BLASTp results have marginal statistical support, the workflow transitions to protein LLM analysis, leveraging their strength in identifying distant patterns and functional relationships. Cases where the two methods yield conflicting predictions represent particularly interesting targets for experimental validation, as they may indicate novel functions or protein families [1] [2].
BLASTp maintains its foundational role in bioinformatics workflows due to its proven accuracy, computational efficiency, and interpretable results for sequences with database homologs. However, the rising capabilities of protein language models demonstrate that AI-driven approaches now offer complementary functionality, particularly for annotating distant homologs and proteins with novel folds [1] [3].
Future directions in protein sequence analysis point toward hybrid frameworks that strategically deploy alignment-based and AI-based methods according to their strengths. The forthcoming transition of BLASTp's default database to ClusteredNR [4] represents an evolution of the traditional paradigm, reducing redundancy while expanding taxonomic coverage. Meanwhile, protein language models continue to advance in their capacity to capture the complex biophysical and evolutionary patterns underlying protein function [3].
For researchers in genomics, drug discovery, and synthetic biology, this evolving landscape offers an expanded toolkit for protein function prediction. Mastering the BLASTp workflow—while understanding its relationship to emerging computational methods—remains essential for rigorous bioinformatics analysis in the coming years.
The accurate annotation of protein function is a cornerstone of modern biology, directly impacting drug discovery, metabolic engineering, and our understanding of disease mechanisms. For decades, homology-based search tools like BLASTp have served as the gold standard for transferring functional knowledge from characterized proteins to novel sequences [1]. This paradigm, however, rests on the availability of evolutionarily related proteins with significant sequence similarity, creating a substantial annotation gap for remote homologs and orphan sequences.
The convergence of protein language models (pLMs) and deep learning (DL) classifiers represents a transformative shift in this landscape. pLMs, pre-trained on millions of protein sequences, learn fundamental principles of protein grammar and generate rich, numerical embeddings that encapsulate structural and functional information [3] [26]. When these embeddings are used as features for specialized DL classifiers, they enable a powerful, alignment-free approach to function prediction that can uncover relationships invisible to traditional sequence comparison methods [27] [28].
Framed within the broader thesis of pLM versus BLASTp annotation research, this guide provides a performance comparison of these integrated pLM-DL pipelines against established benchmarks. We synthesize recent experimental data, detail core methodologies, and provide resources to help researchers and drug development professionals select the optimal tool for their annotation challenges.
Performance Comparison: pLM-DL Models vs. Traditional Methods
Direct comparisons reveal the distinct performance profiles of traditional sequence search, pLM-based, and structure-based methods. The following tables summarize key quantitative findings from recent large-scale benchmarks.
Table 1: Overall Performance Comparison on Enzyme Commission (EC) Number Prediction
| Method | Type | Key Metric | Performance | Reference |
|---|---|---|---|---|
| BLASTp | Sequence Alignment | Overall Accuracy | Marginally Best | [1] |
| ESM2 (with DNN) | pLM + DL | Overall Accuracy | Very High, Complementary to BLASTp | [1] |
| ProtBERT (with DNN) | pLM + DL | Overall Accuracy | Very High | [1] |
| ESM1b (with DNN) | pLM + DL | Overall Accuracy | Very High | [1] |
| One-Hot Encoding (with DL) | Traditional DL | Overall Accuracy | Lower than pLM-based Models | [1] |
Table 2: Remote Homology Search Sensitivity (AUROC) on SCOPe40-test Dataset
| Method | Family-Level (AUROC) | Superfamily-Level (AUROC) | Fold-Level (AUROC) | Reference |
|---|---|---|---|---|
| PLMSearch | 0.928 | 0.826 | 0.438 | [27] |
| MMseqs2 | 0.318 | 0.050 | 0.002 | [27] |
| BLASTp | N/A | N/A | N/A | [27] |
| Foldseek (Structure) | Comparable to PLMSearch | Comparable to PLMSearch | Comparable to PLMSearch | [27] |
| Performance Gain (PLMSearch vs. MMseqs2) | 3x | 16x | 219x | [27] |
Table 3: Performance of Specialized pLM-DL Models on Specific Prediction Tasks
| Model | Task | Performance | Reference |
|---|---|---|---|
| NCSP-PLM | Non-Classical Secreted Protein Prediction | Accuracy: 94.12%, Sensitivity: 91.18%, Specificity: 97.06% | [28] |
| Fine-tuned pLMs (ESM2, ProtT5) | Viral Protein Function Prediction | Significant improvement in embedding quality and downstream task performance vs. pre-trained pLMs | [26] |
Experimental Protocols and Methodologies
To ensure reproducibility and provide clarity on how the data in the previous section was generated, this section outlines the standard experimental protocols used in benchmarking studies.
Standard Benchmarking Workflow for EC Number Prediction
A typical experimental protocol for comparing EC number prediction methods, as used in studies like the assessment of protein LLMs, involves several key stages [1]:
Data Curation and Preprocessing:
- Source: Protein sequences and their corresponding EC numbers are extracted from the UniProt Knowledgebase (UniProtKB). Manually reviewed Swiss-Prot entries are often preferred for creating high-confidence benchmark sets.
- De-redundancy: To avoid bias, sequences are clustered at a specific identity threshold (e.g., using UniRef90 clusters), retaining only a single representative sequence per cluster.
- Data Splitting: The dataset is split into training, validation, and independent test sets, ensuring that no proteins in the test set have high sequence similarity to those in the training set. This rigorously tests the model's ability to generalize.
Feature Extraction for pLM-DL Models:
- pLM Embedding Generation: For each protein sequence in the dataset, a pre-trained pLM (e.g., ESM2, ESM1b, ProtBERT) is used to generate a sequence embedding. This is typically done by taking the hidden state representations from the final or penultimate layer of the model, often by averaging across all amino acid positions to create a fixed-length vector per protein.
- Traditional Feature Generation (Baseline): For baseline models, features are generated using one-hot encoding of amino acid sequences or other classical feature extraction methods.
Model Training and Evaluation:
- Classifier Training: The extracted features (pLM embeddings or one-hot encodings) are used to train a deep learning classifier, such as a fully connected Deep Neural Network (DNN). The task is typically framed as a multi-label classification problem.
- Baseline Comparison: The pLM-DL models are compared against the baseline models and against the direct predictions from BLASTp. For BLASTp, a common approach is to transfer the EC number of the top-hit in the database with a defined E-value or identity cutoff.
- Performance Metrics: Models are evaluated on the held-out test set using metrics like AUROC (Area Under the Receiver Operating Characteristic curve), accuracy, precision, recall, and F1-score.
Workflow for Remote Homology Search (PLMSearch)
The PLMSearch method provides a protocol optimized for detecting remote evolutionary relationships [27]:
- Input and Pre-filtering: Query and target protein sequences are provided as input. An optional pre-filtering step (PfamClan) can quickly remove protein pairs that do not share any known protein domain clan.
- Embedding and Similarity Prediction: Protein sequence embeddings are generated using a large pLM (e.g., ESM2). These embeddings are then fed into a specialized Structural Similarity (SS) Predictor, which is a neural network trained to predict the TM-score (a measure of structural similarity) between two proteins based solely on their embeddings.
- Ranking and Output: The query-target protein pairs are ranked based on their predicted similarity score from the SS-predictor. This ranked list forms the primary output of PLMSearch.
- Alignment (Optional): For top-ranked pairs, a final alignment step (PLMAlign) can be invoked to generate a precise sequence alignment and alignment score using the pLM embeddings.
The following diagram illustrates the logical workflow and decision points in a typical pLM-DL benchmarking experiment, integrating the protocols above:
Successful implementation of pLM-DL pipelines relies on a suite of computational tools and datasets. The table below details key resources referenced in the studies covered in this guide.
Table 4: Key Research Reagents and Computational Tools
| Resource Name | Type | Primary Function in Research | Reference |
|---|---|---|---|
| UniProt Knowledgebase (UniProtKB) | Database | Provides the canonical source of protein sequences with high-quality functional annotations (e.g., EC numbers) for model training and testing. | [1] [3] |
| ESM2 (Evolutionary Scale Modeling) | Protein Language Model | A transformer-based pLM used to generate deep contextual embeddings from protein sequences for downstream prediction tasks. Available in multiple sizes (e.g., ESM2-3B, ESM2-15B). | [1] [27] [26] |
| ProtBERT | Protein Language Model | Another powerful transformer-based pLM, pre-trained on UniRef and BFD, used for generating protein sequence embeddings. | [1] [26] |
| ProtT5 | Protein Language Model | A pLM based on the T5 (Text-to-Text Transfer Transformer) architecture, known for producing high-quality sequence representations. | [27] [26] |
| BLASTp | Software Tool | The standard benchmark for sequence alignment-based function prediction; used for comparison and often in ensemble methods. | [1] [27] |
| MMseqs2 | Software Tool | A highly sensitive and fast sequence search tool used as a baseline for comparing remote homology detection performance. | [27] |
| PLMSearch | Software Suite | An integrated method for remote homology search that uses pLM embeddings and a structural similarity predictor, offering a web server. | [27] |
| SCOPe Database | Database | A curated database of protein structural classifications used as a gold-standard benchmark for evaluating fold-level homology detection. | [27] |
The integration of protein language model embeddings with deep learning classifiers has firmly established itself as a powerful and often superior alternative to traditional BLASTp annotation for specific, high-value scenarios. The experimental data demonstrates that while BLASTp retains a marginal overall advantage for routine annotation, pLM-DL models offer unparalleled sensitivity in detecting remote homologs and can achieve state-of-the-art accuracy on specialized prediction tasks like identifying non-classically secreted proteins.
The choice between these paradigms is not merely a binary one. As research progresses, the most effective strategies are likely to be hybrid, leveraging the speed and reliability of BLASTp for clear homologs while deploying the deep semantic power of pLM-DL models for the "dark matter" of the protein universe—sequences with no close relatives, extreme diversity, or from underrepresented biological families. For researchers and drug developers, this expanding toolkit promises to accelerate the functional elucidation of novel targets, ultimately driving innovation in biomedicine and biotechnology.
Enzyme function prediction is a critical task in bioinformatics, with direct implications for understanding cellular metabolism, drug discovery, and synthetic biology. The Enzyme Commission (EC) number system provides a standardized hierarchical framework (with four digits like 1.2.3.4) for classifying enzyme function [29]. This guide compares the performance of traditional sequence alignment tools, protein language models, and emerging hybrid approaches for EC number prediction, providing researchers with data-driven insights for method selection.
The table below summarizes the core performance characteristics of major EC number prediction approaches, highlighting their respective strengths and limitations.
| Method Type | Representative Tools | Key Strengths | Major Limitations |
|---|---|---|---|
| Sequence Alignment | BLASTp, MMseqs2 [2] [27] | High accuracy for enzymes with close homologs [2] [10] | Fails for novel enzymes without homologs; performance drops sharply at low sequence identity [29] [2] |
| Protein Language Models (PLMs) | ESM2, ESM1b, ProtBERT [2] [10] | Effective for remote homology and enzymes without close homologs; excels when sequence identity <25% [2] [10] | Marginally lower overall accuracy than BLASTp; requires substantial computational resources [2] [10] |
| Structure-Based Models | TopEC [30] | High accuracy (F-score: 0.72) by leveraging 3D structural information; robust to fold bias [30] | Dependent on availability of high-quality 3D structures, which can be scarce [29] [30] |
| Multi-Modal/Hybrid Models | MAPred [29] | State-of-the-art performance by integrating sequence and structural (3Di) data; respects EC number hierarchy [29] | Increased model complexity and computational cost [29] |
Accurately determining enzyme function is fundamental for applications ranging from genome annotation to drug design [3] [29]. However, experimental methods for function determination are time-consuming and resource-intensive, creating a massive gap between the number of known protein sequences and those with experimentally validated functions [3]. As of early 2024, out of over 240 million protein sequences in the UniProt database, less than 0.3% have been experimentally annotated [3]. This gap has driven the development of computational methods for automated function prediction, with the core challenge being to infer the correct EC number from an enzyme's amino acid sequence or structure.
Methodologies in Focus
Traditional Workhorse: Sequence Alignment with BLASTp
BLASTp (Basic Local Alignment Search Tool for proteins) operates on the principle of homology. It identifies similar sequences in annotated databases and transfers functional annotations from the best hits [10]. Its methodology is straightforward: a query protein sequence is compared against a reference database of known sequences using pairwise alignment, and EC numbers are assigned based on the highest sequence similarity matches [2] [10].
The Modern Contender: Protein Language Models (PLMs)
Inspired by large language models in NLP, PLMs like ESM2 and ProtBERT are pre-trained on millions of protein sequences in a self-supervised manner [3] [27]. They learn evolutionary patterns and structural constraints embedded in the sequence data.
Typical Experimental Protocol for PLM-based EC Prediction:
- Feature Extraction: A pre-trained PLM (e.g., ESM2) processes the input amino acid sequence, generating a numerical representation (embedding) for the entire protein or its constituent residues [2] [10].
- Classifier Training: These embeddings are used as input features to train a classifier, typically a fully connected neural network, on a large dataset of enzymes with known EC numbers (e.g., from UniProtKB) [2] [10].
- Hierarchical Prediction: The classifier is often trained as a multi-label model to predict all relevant EC digits simultaneously, accounting for promiscuous and multi-functional enzymes [10].
The Integrated Frontier: Multi-Modal and Hybrid Approaches
The latest methods integrate multiple data types to overcome the limitations of single-modality models.
MAPred (Multi-scale multi-modality Autoregressive Predictor) combines protein sequence with 3D structural information represented as 3Di tokens [29]. Its workflow is detailed below.
PLMSearch offers a different hybrid approach, using a PLM to generate deep sequence representations that are used to predict structural similarity (TM-score), enabling highly sensitive remote homology detection that is much faster than structure search tools [27].
Comparative Performance Analysis
Quantitative Benchmarking
The table below presents key performance metrics from recent comparative studies, offering a direct comparison of different methodologies.
| Method | Approach | Dataset | Key Metric | Performance | Reference |
|---|---|---|---|---|---|
| BLASTp | Sequence Alignment | UniProtKB-based | Overall Accuracy | Marginally better than individual PLMs [2] [10] | [2] |
| ESM2 + FCNN | Protein Language Model | UniProtKB-based | Overall Accuracy | Slightly lower than BLASTp, but complementary [2] [10] | [2] |
| TopEC | 3D Graph Neural Network | PDB300 (Fold Split) | F-score | 0.72 [30] | [30] |
| MAPred | Multi-modal (Seq + 3Di) | New-392, Price, New-815 | Accuracy | Outperforms existing models [29] | [29] |
| PLMSearch | PLM-based Similarity | SCOPe40-test (Fold-level) | AUROC | 0.438 (vs. MMseqs2: 0.002) [27] | [27] |
Strengths, Weaknesses, and Complementary Performance
BLASTp vs. PLMs: While BLASTp holds a slight overall edge, the performances are complementary [2] [10]. PLMs like ESM2 demonstrate a significant advantage in predicting functions for remote homologs and enzymes with no close relatives, particularly when sequence identity to known proteins falls below 25% [2] [10]. BLASTp excels when strong homologs exist in the database.
The Impact of Data Modality: Models incorporating structural information (e.g., MAPred, TopEC) consistently achieve superior performance [29] [30]. This is because an enzyme's function is directly determined by its 3D structure and the chemical environment of its active site, which cannot be fully captured by sequence alone.
Successful development and application of EC prediction pipelines rely on several key resources.
| Resource Name | Type | Primary Function in EC Prediction |
|---|---|---|
| UniProt Knowledgebase (UniProtKB) | Database | Provides curated protein sequences and functional annotations (including EC numbers) for model training and validation [3] [10]. |
| Protein Data Bank (PDB) | Database | Repository of experimentally determined protein 3D structures, used for training structure-based models and for ground-truth validation [30]. |
| ESM2/ESM1b | Protein Language Model | Pre-trained models used to convert amino acid sequences into informative numerical embeddings (features) for downstream classification tasks [2] [10] [27]. |
| ProstT5 | Software | Predicts 3Di structural tokens from a protein sequence, enabling the incorporation of structural information without needing a 3D coordinate file [29]. |
| AlphaFold2 | Software | Provides high-accuracy protein structure predictions, expanding the potential application of structure-based function prediction methods [30] [27]. |
The field of EC number prediction is evolving from reliance on single-method approaches to integrated, multi-modal pipelines. While BLASTp remains a robust and widely used tool, protein language models have established themselves as powerful alternatives, especially for challenging cases of remote homology [2] [27]. The most accurate methods now combine sequence and structural information in architectures that respect the hierarchical nature of the EC number system [29].
Future progress will likely be driven by several key trends: the continued scaling of protein language models, the increased availability of high-quality predicted structures, and the development of more sophisticated multi-modal fusion techniques. For researchers, the choice of tool should be guided by the specific task—BLASTp for routine annotations with clear homologs, PLMs for remote homology detection, and hybrid models like MAPred for the most challenging and accuracy-critical applications.
Protein Language Models (pLMs), inspired by breakthroughs in natural language processing, are revolutionizing computational biology. Trained on millions of protein sequences through self-supervised learning, these models learn fundamental principles of protein grammar and semantics, extending their utility far beyond traditional sequence annotation tasks [3]. While tools like BLASTp have served as the gold standard for homology-based function prediction for decades, pLMs offer a transformative approach by capturing complex evolutionary patterns and structural constraints directly from sequences [31] [32].
This paradigm shift enables unprecedented applications in protein engineering and therapeutic design. pLMs can generate functional de novo protein sequences, predict evolutionary trajectories, and guide the optimization of biotherapeutic candidates with efficiency surpassing traditional methods [33] [32]. This guide provides a comprehensive comparison of pLM capabilities against established methods, detailing performance metrics, experimental protocols, and practical workflows to inform researchers in biotechnology and drug development.
Performance Comparison: pLMs vs. Traditional Methods
1Function Prediction and Remote Homology Detection
pLMs significantly enhance sensitivity in detecting remote homologies where sequence identity is low, a area where traditional alignment-based methods struggle.
Table 1: Comparative Performance in Enzyme Commission (EC) Number Prediction
| Method | Overall Accuracy | Performance on Sequences with <25% Identity | Key Strengths |
|---|---|---|---|
| BLASTp | Marginally better overall [1] | Lower performance | excels when clear homologs exist in databases [1] |
| ESM2 (pLM) | High, complementary to BLASTp [1] | More accurate predictions [1] | Better for difficult-to-annotate enzymes without close homologs [1] |
| ESM1b (pLM) | High [1] | Good | Effective feature extraction for function prediction [1] |
| ProtBERT (pLM) | High [1] | Good | Can be fine-tuned for specific prediction tasks [1] |
| Combined BLASTp + pLM | Surpasses individual techniques [1] | High | Leverages complementary strengths for maximum coverage [1] |
For remote homology search, specialized pLM-powered tools like PLMSearch demonstrate exceptional performance. In benchmarks searching millions of protein pairs, PLMSearch operated in seconds like MMseqs2 but increased sensitivity by more than threefold, recalling most remote homology pairs with dissimilar sequences but similar structures [27]. Its performance was comparable to state-of-the-art structure search methods while requiring only sequence inputs [27].
Table 2: Performance in Remote Homology Search (SCOPe40-test dataset)
| Method | Family-level AUROC | Superfamily-level AUROC | Fold-level AUROC | Search Speed (4.8M pairs) |
|---|---|---|---|---|
| MMseqs2 | 0.318 | 0.050 | 0.002 | Seconds [27] |
| PLMSearch (ESM-based) | 0.928 | 0.826 | 0.438 | 4 seconds [27] |
| TM-align (Structure-based) | High (Not Shown) | High (Not Shown) | High (Not Shown) | 11,303 seconds [27] |
2Protein Design and Engineering
pLMs have demonstrated remarkable success in generating novel, functional proteins, a task beyond the scope of tools like BLASTp.
- De Novo Generation: Models like ProtGPT2 can generate de novo protein sequences that populate "dark" areas of the proteome, regions where protein structures have never been observed in nature [34]. Similarly, ProGen, a language model trained on 280 million protein sequences across 19,000 families, can generate functional protein sequences with predictable functions [33].
- Experimental Validation: When ProGen was fine-tuned on lysozyme families, the resulting artificial proteins exhibited similar catalytic efficiencies to natural lysozymes, despite having sequence identities as low as 31.4% to any known natural protein [33]. This demonstrates pLMs' ability to capture the essential functional determinants beyond mere sequence similarity.
- Therapeutic Antibody Engineering: pLMs can guide protein evolution without task-specific training. By selecting mutations with higher language-model likelihood than wild-type (positive "evolutionary velocity"), researchers have achieved highly efficient, machine-learning-guided antibody affinity maturation against diverse viral antigens [32]. This process occurred without providing the model with knowledge of the antigen, protein structure, or task-specific data [32].
Experimental Protocols and Workflows
1Protocol: Enzyme Function Prediction with pLMs
This protocol outlines the methodology for predicting Enzyme Commission (EC) numbers using pLMs as feature extractors, based on the comparative assessment by [1].
Data Preparation:
- Extract protein sequences and their EC numbers from UniProtKB (Swiss-Prot and TrEMBL).
- Use only UniRef90 cluster representatives to reduce sequence redundancy.
- Format the prediction as a multi-label classification problem. For an enzyme with multiple EC numbers (e.g., 1.1.1.1 and 4.1.1.1), the label vector includes all hierarchical levels (1, 1.1, 1.1.1, 1.1.1.1, 4, 4.1, 4.1.1, 4.1.1.1).
Feature Extraction:
- Obtain sequence embeddings (numeric representations) from a pre-trained pLM.
- Pass each protein sequence through a model like ESM2, ESM1b, or ProtBERT.
- Extract the hidden layer representations (typically from the final or penultimate layer) for each amino acid position.
- Apply a pooling operation (e.g., mean pooling) across the sequence length to create a fixed-length, global protein embedding.
- Obtain sequence embeddings (numeric representations) from a pre-trained pLM.
Model Training and Prediction:
- Use the extracted protein embeddings as input features for a supervised classifier, typically a fully connected neural network.
- Train the model to predict the binary label vectors, learning the association between the pLM-derived features and the hierarchical EC numbers.
- The trained model can then predict EC numbers for novel, unannotated protein sequences based on their pLM embeddings alone.
Figure 1: Workflow for pLM-based enzyme function prediction
2Protocol: pLM-Guided Therapeutic Protein Engineering
This protocol describes the use of structure-informed pLMs for antibody engineering, as validated in [32].
Model Selection and Input:
- Utilize a structure-informed protein language model. This model incorporates protein backbone coordinates (without side-chain information) to learn general principles of binding that generalize across protein complexes.
- Input the sequence (and/or structural data, if required) of the therapeutic protein candidate (e.g., a wild-type antibody).
In Silico Mutagenesis and Scoring:
- Generate a library of possible single-point mutants (in silico) of the parent sequence.
- Score each variant by calculating its "likelihood" or "evolutionary velocity" using the pLM. Mutants with a higher likelihood than the wild-type are predicted to have positive evolutionary momentum and potentially improved fitness.
- Rank all generated variants based on their model scores.
Experimental Validation:
- Synthesize the top-ranked candidate sequences (e.g., the top 10-50) for laboratory testing.
- Express the proteins and assay for the desired property (e.g., binding affinity, neutralization potency, thermal stability).
- The best-performing variants from this first round can be used as new parent sequences for iterative rounds of optimization, following the same protocol.
Figure 2: Workflow for pLM-guided therapeutic antibody engineering
The Scientist's Toolkit: Key Research Reagents and Solutions
Table 3: Essential Resources for pLM Research in Protein Engineering
| Resource/Solution | Type | Primary Function in Research |
|---|---|---|
| UniProtKB Database [1] | Database | Comprehensive, high-quality protein sequence and functional annotation data for model training and validation. |
| ESM2 Model Suite [1] [31] | Pre-trained pLM | Provides state-of-the-art sequence embeddings for function prediction and is a backbone for developing specialized tools. |
| ProGen [33] | Pre-trained pLM | Conditional generation of novel, functional protein sequences across diverse protein families. |
| PLMSearch Tool [27] | Software | Fast, sensitive remote homology search using pLM embeddings, bridging sequence and structure search sensitivity. |
| Structure-Informed pLM [32] | Computational Model | Incorporates structural data to improve predictions of fitness and binding for therapeutic protein engineering. |
| AlphaFold DB [3] | Database | Repository of predicted protein structures; provides structural context and validation for pLM-based predictions and designs. |
Protein Language Models have unequivocally transcended their initial role as annotation assistants to BLASTp. While BLASTp retains value for routine homology detection, pLMs offer a fundamental advance by enabling the sensitive detection of remote homologies, the rational design of novel functional proteins, and the accelerated engineering of biotherapeutics. The experimental data confirms that pLMs are not merely incremental improvements but represent a paradigm shift, providing a powerful, unified approach to extract functional, evolutionary, and structural insights from sequence alone. For researchers in drug development and protein engineering, integrating these tools into existing workflows is becoming essential for maintaining a competitive edge.
Protein Language Models (pLMs) represent a transformative shift in computational biology, offering a powerful alternative to traditional sequence alignment tools like BLASTp for identifying novel drug targets. This guide provides an objective comparison of these methodologies, focusing on their performance in key drug discovery applications. While BLASTp remains a robust tool for identifying close homologs, advanced pLMs demonstrate superior capabilities in predicting protein functions, interactions, and functional sites—particularly for proteins with distant evolutionary relationships or no known homologs. The integration of pLMs into target identification workflows is now enabling researchers to uncover previously inaccessible therapeutic opportunities with greater efficiency and accuracy.
Performance Benchmarking: pLMs vs. BLASTp
Core Functional Annotation Performance
Table 1: Comparison of pLMs and BLASTp on Key Annotation Tasks
| Annotation Task | Metric | BLASTp | pLM-Based Model | Improvement | Context |
|---|---|---|---|---|---|
| Enzyme Commission (EC) Number Prediction | Overall Accuracy | Benchmark | ESM2 + DNN | Slightly lower than BLASTp [10] | General enzyme function prediction |
| EC Number Prediction (Low Homology) | Accuracy on sequences <25% identity | Lower | ESM2 + DNN | Significantly higher [10] | Difficult cases without clear homologs |
| Protein Family (Pfam) Annotation | Classification Error | Benchmark | ESM/ProtTrans Embeddings | 60% reduction [35] | Protein domain annotation |
| Active Site Annotation | Recall | Benchmark | EasIFA (pLM+Structure) | 7.57% increase [36] | Catalytic site identification for drug targeting |
| Active Site Annotation | Speed (Inference) | Benchmark | EasIFA (pLM+Structure) | 10x faster [36] | High-throughput screening applications |
Protein-Protein Interaction (PPI) Prediction Performance
Table 2: Cross-Species PPI Prediction Performance (AUPR)
| Test Species | BLAST-Derived Methods | PLM-Interact | Improvement vs. Second Best | Training Data |
|---|---|---|---|---|
| Mouse | 0.84 (TUnA) [16] | 0.86 | 2% [16] | Human PPIs |
| Fly | 0.76 (TUnA) [16] | 0.82 | 8% [16] | Human PPIs |
| Worm | 0.74 (TUnA) [16] | 0.79 | 6% [16] | Human PPIs |
| Yeast | 0.64 (TUnA) [16] | 0.71 | 10% [16] | Human PPIs |
| E. coli | 0.68 (TUnA) [16] | 0.72 | 7% [16] | Human PPIs |
Experimental Protocols for pLM Validation
Protocol: PLM-Interact for Protein-Protein Interaction Prediction
Application in Drug Discovery: Identifying novel protein-protein interactions as potential therapeutic targets, particularly for disrupting pathogenic pathways.
Detailed Methodology:
- Model Architecture: Based on ESM-2 transformer architecture with two key extensions:
- Increased permissible sequence length to accommodate paired protein sequences
- Implementation of "next sentence prediction" task fine-tuned on interacting/non-interacting pairs [16]
Training Strategy:
- Pre-trained on human PPI data (421,792 protein pairs: 38,344 positive, 383,448 negative)
- Balanced loss function with 1:10 ratio between classification loss and masked language modeling loss
- Joint encoding of protein pairs enables attention mechanisms to learn inter-protein residue associations [16]
Validation Framework:
- Cross-species validation trained on human data, tested on mouse, fly, worm, yeast, and E. coli
- Benchmarking against six established PPI prediction methods (TUnA, TT3D, Topsy-Turvy, D-SCRIPT, PIPR, DeepPPI)
- Evaluation using Area Under Precision-Rcall Curve (AUPR) to account for class imbalance [16]
Mutation Effect Prediction:
- Fine-tuned on IntAct mutation data (6,979 annotated mutation effects)
- Predicts impact of mutations on interaction strength (increasing or decreasing) [16]
Protocol: EasIFA for Enzyme Active Site Annotation
Application in Drug Discovery: Precise identification of enzyme active sites for inhibitor design and allosteric drug development.
Detailed Methodology:
- Multi-Modal Architecture:
- Sequence Representation: ESM-2 embeddings capturing evolutionary information
- Structural Representation: 3D structural encoder processing spatial coordinates
- Reaction Information: Biochemical reaction data integrated via cross-attention mechanisms [36]
Feature Fusion:
- Implements multi-modal cross-attention framework to align protein-level information with enzymatic reaction knowledge
- Combines latent enzyme representations from pLM and structural encoder
- Self-supervised pre-training on organic chemical reaction databases [36]
Training Data:
- Leverages both coarse-annotated enzyme databases and high-precision manually curated datasets
- Transfer learning capability between databases with different annotation standards [36]
Validation:
- Benchmarking against BLASTp and structure-based methods (AEGAN, SiteMap)
- Evaluation metrics: Recall, Precision, F1-score, Matthews Correlation Coefficient (MCC)
- Speed assessment compared to PSSM-based methods [36]
Protocol: Contrastive Learning for Effector Prediction (CLEF)
Application in Drug Discovery: Identification of bacterial virulence factors as targets for anti-infective therapies.
Detailed Methodology:
- Dual-Encoder Architecture:
- Encoder A: Processes ESM-2 representations through transformer layers
- Encoder B: Projects biological modality features through MLP layers [37]
Contrastive Learning Framework:
- Pre-training with InfoNCE loss to align PLM representations with biological features
- Integration of multiple modalities: Secretion Embedding, Annotation Text, 3D structural features
- Generates coordinated cross-modality representations in latent space [37]
Biological Feature Integration:
- Secretion Embedding: Classification features from secretion effectors
- Annotation Text: GO terms and protein annotations encoded via BioBERT
- 3Di Features: Structural token sequences from Foldseek spatial information [37]
Validation:
- Testing on enteric pathogens (E. coli, Salmonella Typhimurium, Edwardsiella piscicida)
- Experimental validation of predicted T3SEs and T6SEs
- Clustering performance metrics: ARI, NMI, ASW [37]
Visualization of Key Methodological Relationships
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Research Tools for pLM-Based Target Identification
| Tool/Category | Specific Examples | Function in Research | Application Context |
|---|---|---|---|
| Pre-trained pLMs | ESM-2, ProtTrans, ProtBERT | Generate protein sequence embeddings capturing evolutionary and structural information [35] [10] | Base feature extraction for all downstream tasks |
| Structural Databases | PDB, AFDB (AlphaFold DB) | Provide experimental and predicted structures for multi-modal integration [37] | Structure-aware function prediction |
| Interaction Databases | IntAct, BioGRID, STRING | Source of validated PPIs for model training and benchmarking [16] [38] [39] | PPI prediction and validation |
| Functional Annotations | UniProt, Pfam, Gene Ontology | Gold-standard labels for model training and performance evaluation [3] [35] | Functional prediction benchmarks |
| Specialized Software | Foldseek, ProstT5 | Encode structural information into machine-readable features [37] | Structural modality integration |
| Experimental Validation | Yeast two-hybrid, TAP-MS, PROPER-seq | Experimental methods for validating computational predictions [38] [39] | Ground truth establishment |
The comprehensive benchmarking data presented in this guide demonstrates that while BLASTp maintains advantages for straightforward homology-based annotation, advanced pLMs and their multi-modal extensions offer significant improvements for the most challenging and clinically relevant target identification tasks. The key differentiators for pLMs include their ability to function effectively with low-homology proteins, integrate diverse biological data types, and predict complex molecular interactions with state-of-the-art accuracy.
For drug discovery pipelines, we recommend a hybrid approach: utilizing BLASTp for initial sequence characterization while employing pLM-based methods for identifying functional sites, predicting interactions, and characterizing proteins of unknown function. As pLMs continue to evolve, their capacity to integrate structural, chemical, and multi-omic data will likely establish them as the universal foundation for computational target identification in pharmaceutical research.
Beyond the Basics: Troubleshooting Limitations and Optimizing Your Annotation Pipeline
This guide provides an objective comparison between traditional homology-based tools like BLASTp and emerging Protein Language Models (pLMs) for enzyme function prediction. As the volume of unannotated protein sequences grows exponentially—with less than 0.3% of sequences in UniProt having experimentally validated functions—the limitation of conventional methods that rely on sequence similarity has become a critical research bottleneck [3]. Our analysis, based on recent comparative studies, reveals that while BLASTp maintains a marginal overall performance advantage, pLMs demonstrate superior capability in predicting functions for non-homologous enzymes and those with sequence identity below 25% [1] [2]. This complementary performance profile suggests that an integrated approach leveraging both technologies represents the future of comprehensive enzyme annotation.
Performance Benchmarking: pLMs vs. BLASTp
Quantitative Performance Comparison
Table 1: Overall performance comparison between BLASTp and protein language models for EC number prediction
| Method | Overall Accuracy | Performance on Low-Homology Sequences (<25% identity) | Key Strengths | Major Limitations |
|---|---|---|---|---|
| BLASTp | Marginally superior in direct comparisons [1] | Significant performance degradation | Excellent for closely related sequences with clear homologs | Cannot annotate enzymes without homologous sequences in databases [1] |
| Protein Language Models (ESM2) | Slightly lower overall but complementary to BLASTp [1] | Superior performance on difficult-to-annotate enzymes [1] [2] | Effective for remote homology detection and non-homologous enzymes [1] | Still requires improvement to become gold standard for mainstream annotation [1] |
| Combined Approach | Performance exceeds individual methods [1] | Comprehensive coverage across homology spectrum | Leverages strengths of both methodologies | More complex implementation pipeline |
Specialized Performance Insights
Table 2: Performance characteristics of individual protein language models
| pLM Model | Relative Performance | Key Applications | Architecture |
|---|---|---|---|
| ESM2 | Best performer among pLMs tested [1] [2] | EC number prediction, protein-protein interactions [16] | Transformer-based [1] |
| ESM1b | Competitive alternative to ESM2 [1] [3] | Protein function prediction, structure prediction [3] | Transformer-based [1] |
| ProtBERT | Effective for EC number prediction [1] | Enzyme function prediction [1] | BERT-style transformer [1] |
| PLM-interact | State-of-the-art for PPI prediction [16] | Protein-protein interactions, mutation effects [16] | Fine-tuned ESM-2 with next-sentence prediction [16] |
Experimental Protocols and Methodologies
Standardized Evaluation Framework
Recent comparative studies have established rigorous experimental protocols to evaluate BLASTp against protein language models objectively. The key methodological considerations include:
Dataset Construction: Studies utilized UniProtKB data (SwissProt and TrEMBL) downloaded in February 2023, processing only UniRef90 cluster representatives to enhance dataset quality and reduce redundancy [1]. This approach selects representatives based on entry quality, annotation score, organism relevance, and sequence length.
Problem Formulation: EC number prediction was defined as a multi-label classification problem incorporating promiscuous and multi-functional enzymes. The global approach for hierarchical multi-label classification challenges a single classifier to predict the entire hierarchy of labels and their relationships [1].
Embedding Generation: For pLM-based approaches, models including ESM2, ESM1b, and ProtBERT were used as feature extractors. The embeddings from these models were then combined with fully connected neural networks for EC number prediction [1].
Critical Workflow Components
BLASTp Implementation: Standard protein BLAST (BLASTp) programs search protein databases using a protein query, with parameter values that differ from defaults specifically highlighted in evaluation interfaces [40].
PLM-interact Methodology: This extended pLM approach implements two key innovations: (1) longer permissible sequence lengths in paired masked-language training to accommodate residues from both proteins, and (2) implementation of "next sentence" prediction to fine-tune all layers of ESM-2 where the model is trained with a binary label indicating interaction status [16].
Diagram 1: Integrated workflow for enzyme function annotation combining BLASTp and pLM approaches
Table 3: Key research reagents and computational resources for enzyme function prediction
| Resource | Type | Primary Function | Access |
|---|---|---|---|
| UniProtKB | Database | Comprehensive protein sequence and functional information [1] | Publicly available |
| ESM-2 | Protein Language Model | Generate embeddings for function prediction [1] [16] | Open source |
| ProtBERT | Protein Language Model | BERT-based protein sequence representations [1] | Open source |
| StarPDB | Annotation Server | Structural annotation based on PDB similarity [41] | Web server |
| FoldSeek | Structure Search | Fast protein structure similarity search [42] | Open source |
| PLM-interact | Specialized pLM | Protein-protein interaction prediction [16] | Research implementation |
Technical Implementation Guide
pLM Integration Architecture
The most effective implementations of pLMs for enzyme function prediction follow a structured architectural pattern:
Embedding Extraction: Protein sequences are processed through pre-trained pLMs (ESM2, ProtBERT) to generate fixed-size vector representations (embeddings) that encapsulate evolutionary, structural, and functional information [1] [43].
Classification Head: These embeddings are then fed into fully connected neural networks specifically trained for EC number prediction. This approach has been shown to surpass the performance of deep learning models relying on one-hot encodings of amino acid sequences [1].
Hierarchical Prediction: The multi-label classification framework addresses the EC number hierarchy comprehensively, predicting all relevant levels (e.g., for EC 1.1.1.1, the model predicts 1, 1.1, 1.1.1, and 1.1.1.1) [1].
Hybrid Deployment Strategy
Diagram 2: Decision workflow for hybrid BLASTp-pLM enzyme annotation system
Future Directions and Research Opportunities
The rapid evolution of protein language models suggests several promising research directions:
Embedding-Enhanced Search: Integrating pLM-derived embeddings with traditional sequence databases could extend homology detection beyond sequence similarity to functional similarity [42].
Multi-Modal Approaches: Combining sequence embeddings with predicted structural information from tools like AlphaFold could address scenarios where both sequence similarity is low and structural similarity is moderate [42].
Specialized Fine-Tuning: Domain-specific fine-tuning of general pLMs for particular enzyme classes or families may further enhance performance for challenging annotation tasks [16].
The comparative assessment between BLASTp and protein language models reveals a nuanced landscape for enzyme function prediction. While BLASTp remains slightly superior for routine annotation tasks involving enzymes with clear homologs, protein language models excel precisely where BLASTp encounters fundamental limitations—particularly for non-homologous enzymes and sequences with less than 25% identity to characterized proteins [1] [2].
This complementary relationship underscores that these technologies are not mutually exclusive but rather function most effectively when integrated. As protein language models continue to evolve, they are poised to address the critical "blind spots" in homology-based approaches, ultimately enabling more comprehensive annotation of the rapidly expanding universe of protein sequences. For research teams seeking to maximize annotation coverage, a hybrid implementation that strategically deploys both BLASTp and pLMs based on sequence characteristics represents the current state-of-the-art approach.
In the field of bioinformatics, detecting homologous relationships and annotating protein function have long been dominated by alignment-based methods like BLASTp. These tools operate on the principle that sequence similarity implies functional similarity and evolutionary relatedness. However, this approach encounters a fundamental limitation: as evolutionary distance increases, sequences diverge to the point where their shared ancestry is no longer detectable through direct sequence comparison. This creates a significant "twilight zone" where remote homologs with similar structures and functions exhibit sequence identity below 25%, confounding traditional search methods [1] [27].
The emergence of protein Language Models (pLMs) represents a paradigm shift in computational biology. Trained on millions of protein sequences through self-supervised learning, pLMs learn the underlying "grammar" of protein sequences, capturing complex evolutionary patterns, structural constraints, and functional motifs that extend beyond simple amino acid identity [14] [9]. This capability enables pLMs to identify distant evolutionary relationships that traditional methods miss, offering new opportunities for protein annotation, functional prediction, and therapeutic discovery.
This guide provides an objective comparison of pLM and BLASTp performance for low-identity protein annotation, presenting experimental data and methodologies to help researchers select appropriate tools for their specific challenges.
Performance Comparison: pLMs vs. BLASTp at Low Sequence Identity
Quantitative Benchmarking Across Critical Tasks
Table 1: Performance comparison of pLMs and BLASTp on low-identity protein annotation tasks.
| Annotation Task | Metric | BLASTp Performance | pLM Performance | Improvement | Notes |
|---|---|---|---|---|---|
| Remote Homology Detection (SCOPe40-test) | AUROC (Fold-level) | 0.002 (MMseqs2) | 0.438 (PLMSearch) | 219x | PLMSearch compared to sequence search method [27] |
| Enzyme Commission Number Prediction | Accuracy (<25% identity) | High with homologs | ESM2 excels without homologs | Complementary | BLASTp marginally better overall; ESM2 better for difficult cases [1] |
| Intrinsically Disordered Region Prediction | Accuracy | N/A | Superior with FusionEncoder | Significant | FusionEncoder integrates traditional features with pLM embeddings [44] |
| DNA-Binding Protein Identification | Performance on remote homologs | Baseline | Enhanced by ensemble methods | Improved | Combining top tools with BLAST improves capability [45] |
Key Performance Insights
Complementary Strengths: While BLASTp maintains an advantage when closely related sequences exist in databases, pLMs demonstrate superior capability for remote homology detection and annotation of orphan sequences with no clear homologs [1] [27].
Sensitivity vs. Specificity Trade-offs: pLMs like PLMSearch achieve fold-level sensitivity improvements of over 200x compared to sequence-based methods while maintaining search speeds comparable to fast tools like MMseqs2 [27].
Functional Prediction Advantages: For structured functional classification such as Enzyme Commission (EC) numbers, pLMs provide more accurate predictions for enzymes where sequence identity to characterized proteins falls below the 25% threshold [1].
Experimental Protocols and Methodologies
Standardized Evaluation Frameworks
To ensure fair comparison between pLMs and alignment-based methods, researchers have established rigorous benchmarking protocols:
3.1.1 Homology Detection Evaluation
- Dataset: SCOPe40-test (2,207 proteins) with all-versus-all search (4.87 million pairs) [27]
- Metrics: Area Under Receiver Operating Characteristic (AUROC) at family, superfamily, and fold levels
- Ground Truth: Structural similarity (TM-score) as reference [27]
- Methods Compared: PLMSearch (pLM-based) vs. MMseqs2, BLASTp, HHblits (alignment-based)
3.1.2 Enzyme Function Prediction Protocol
- Task Definition: Multi-label EC number classification incorporating promiscuous and multi-functional enzymes [1]
- Data Processing: UniProtKB sequences clustered with UniRef90 at 25% identity threshold [1]
- Evaluation Focus: Performance on sequences with <25% identity to training data [1]
- pLM Architecture: ESM2 embeddings fed into fully connected neural networks [1]
pLM-Specific Training and Fine-Tuning Approaches
Domain Adaptation for Underrepresented Proteins: Fine-tuning general pLMs on viral protein sequences using Low-Rank Adaptation (LoRA) significantly enhances representation quality for these underrepresented "dark matter" proteins, demonstrating the adaptability of pLMs to specific biological domains [26].
Multi-Semantic Feature Integration: FusionEncoder employs an LSTM-based fusion network to integrate traditional biological features (evolutionary profiles, physicochemical properties) with pLM embeddings, demonstrating that hybrid approaches outperform single-modality models for challenging tasks like intrinsically disordered region prediction [44].
Technical Breakdown: How pLMs Overcome the Low-Identity Barrier
Architectural Advantages Over Sequence Alignment
Table 2: Fundamental differences between BLASTp and pLM approaches.
| Feature | BLASTp (Alignment-Based) | Protein Language Models |
|---|---|---|
| Core Principle | Local/global sequence alignment | Contextual semantic understanding |
| Evolutionary Signal | Direct residue conservation | Latent evolutionary patterns |
| Context Awareness | Limited to alignment region | Whole-sequence context via self-attention |
| Information Utilization | Explicit sequence similarity | Implicit structural/functional constraints |
| Dependency | Database content and quality | Training data distribution and diversity |
| Strengths | High precision with close homologs | Remote homology detection, orphan sequence annotation |
The Representation Learning Advantage
pLMs generate embedding vectors that encapsulate rich biological information including evolutionary relationships, structural features, and functional properties [26] [14]. These embeddings form a continuous semantic space where proteins with similar functions or structures cluster together regardless of sequence similarity. This enables identification of distant relationships through geometric proximity in embedding space rather than direct sequence alignment [27].
Figure 1: Comparison of BLASTp alignment-based approach versus pLM semantic understanding approach for protein analysis.
Research Reagent Solutions: Essential Tools for Low-Identity Protein Annotation
Table 3: Key computational tools and resources for low-identity protein annotation.
| Tool Name | Type | Primary Function | Advantages for Low-Identity Tasks |
|---|---|---|---|
| PLMSearch | pLM-based search | Remote homology detection | 3-219x sensitivity improvement over MMseqs2; structural search-like performance [27] |
| ESM-2 | Protein language model | Feature extraction/embedding | Best-performing pLM for EC number prediction; excels without homologs [1] |
| FusionEncoder | Hybrid prediction | Disordered region identification | Integrates traditional features with pLM embeddings; superior accuracy [44] |
| ProtT5 | Protein language model | Feature extraction | Used in embedding-based annotation transfer (EAT) [27] |
| pLM-BLAST | pLM-enhanced alignment | Homology detection | Combines pLM representations with Smith-Waterman algorithm [27] |
The experimental evidence demonstrates that protein language models have established a definitive advantage over alignment-based methods like BLASTp for protein annotation in the low-identity regime (<25% sequence identity). pLMs fundamentally overcome the limitations of direct sequence comparison by learning the deep semantic patterns and evolutionary constraints that persist even when sequences have diverged beyond recognition by traditional methods.
However, this does not render BLASTp obsolete. Rather, the two approaches serve complementary roles: BLASTp remains highly effective and efficient for detecting close homologs and establishing clear evolutionary relationships, while pLMs excel at identifying distant evolutionary connections and annotating proteins without clear homologs [1] [27]. The most effective annotation pipelines increasingly combine both approaches, leveraging their respective strengths to achieve comprehensive protein characterization across the entire evolutionary spectrum.
For researchers and drug development professionals, this expanded toolkit enables more complete proteome annotation, better identification of distant disease homologs across species, and new opportunities for discovering previously unrecognized protein functions and interactions—ultimately accelerating therapeutic discovery and biological understanding.
The application of Large Language Models (Sci-LLMs) to biological discovery faces a fundamental preprocessing challenge: the tokenization dilemma. This refers to the inherent difficulty in converting raw biomolecular sequences into discrete tokens that LLMs can process effectively. Whether treating sequences as a specialized language or a separate modality, current strategies fundamentally limit model performance by either fragmenting functional motifs or creating formidable alignment challenges [46]. Scientific LLMs attempting to interpret low-level sequence data directly often struggle with the informational noise present in raw sequences, hindering their reasoning capacity on complex biological tasks [46].
Within protein function prediction, this challenge manifests in the ongoing comparison between traditional similarity search tools like BLASTp and emerging protein language models (PLMs). BLASTp leverages evolutionary relationships through sequence alignment, while PLMs like ESM2 and ProtBERT utilize self-supervised learning on massive protein sequence datasets to capture deeper semantic and structural patterns [2] [1] [9]. This article systematically evaluates how providing high-level structured context—rather than raw sequences—represents a paradigm shift that unlocks the true reasoning potential of Sci-LLMs, positioning them not as sequence decoders but as powerful engines for synthesizing established biological knowledge [46].
Experimental Protocols: Assessing Contextual Input Strategies
Experimental Design for Input Modality Comparison
To quantitatively assess the tokenization dilemma and contextual approach, researchers have implemented systematic experimental frameworks comparing three distinct input strategies across biological reasoning tasks [46]:
- Sequence-only mode: Raw biomolecular sequences are provided directly to Sci-LLMs using specialized tokenization schemes.
- Context-only mode: High-level structured information derived from bioinformatics tools (e.g., functional domains, structural features, evolutionary profiles) replaces raw sequences.
- Combined mode: Both raw sequences and their contextual representations are provided to the models.
The performance evaluation employs standard metrics including area under the receiver operating characteristic curve (AUROC), precision-recall curves, and accuracy rates across different hierarchy levels of enzyme classification (family, superfamily, fold) [2] [1]. Benchmarking occurs on carefully curated datasets like SCOPe40-test and Swiss-Prot after filtering homologs from training data to ensure rigorous evaluation [27].
Protein Language Model Evaluation Protocol
Comparative assessments of protein LLMs versus traditional methods follow established experimental protocols [2] [1]:
- Data Collection: SwissProt and TrEMBL protein data with UniRef90 cluster representatives are extracted from UniProtKB.
- Feature Extraction: For PLMs (ESM2, ESM1b, ProtBERT), embeddings are generated as sequence representations. For traditional methods, one-hot encodings or alignment scores are used.
- Model Training: Fully connected neural networks are trained on PLM embeddings, compared with DeepEC and D-SPACE models using one-hot encodings.
- BLASTp Baseline: BLASTp searches are conducted against reference databases with standard parameters.
- Evaluation: Performance is measured through multi-label classification accuracy at different EC number hierarchy levels, with special attention to low-identity sequences (<25% identity).
Figure 1: Experimental workflow for comparing input modalities in Sci-LLMs.
Results: Contextual Input Outperforms Sequence-Based Approaches
Quantitative Comparison of Input Modalities
Striking results from systematic comparisons reveal that the context-only approach consistently and substantially outperforms all other input modes across biological reasoning tasks [46]. Even more revealing, the inclusion of raw sequences alongside their high-level contextual representations consistently degrades performance, indicating that raw sequences act as informational noise even for models with specialized tokenization schemes [46].
Table 1: Performance comparison of input modalities for Sci-LLMs
| Input Modality | Performance Level | Advantages | Limitations |
|---|---|---|---|
| Sequence-only | Substantially lower | Direct sequence processing | Loses functional motif information |
| Context-only | Consistently superior | Leverages established bioinformatics knowledge | Depends on quality of context sources |
| Combined | Degraded performance | Comprehensive input | Raw sequences add informational noise |
Performance Comparison: PLMs vs. BLASTp
Recent comprehensive evaluations illuminate the complementary strengths of protein language models and traditional BLASTp for enzyme commission number prediction [2] [1]. While BLASTp provides marginally better results overall, deep learning models using PLM embeddings complement BLASTp's capabilities, with each approach excelling in different scenarios.
Table 2: Performance comparison of protein LLMs and BLASTp for EC number prediction
| Method | Overall Accuracy | Performance on Low-Identity Sequences (<25%) | Key Strengths |
|---|---|---|---|
| BLASTp | Marginally better | Limited performance | Excellent for sequences with clear homologs |
| ESM2 | High | Superior performance | Accurate predictions for difficult-to-annotate enzymes |
| ESM1b | Moderate | Good performance | Balanced approach |
| ProtBERT | Moderate | Good performance | Alternative architecture |
| ESM2 + BLASTp | Highest combined | Comprehensive coverage | Complementary strengths |
The ESM2 model emerged as the most effective among tested PLMs, providing more accurate predictions on difficult annotation tasks and for enzymes without close homologs [2]. Crucially, these studies demonstrate that PLMs still require improvement to become the gold standard over BLASTp in mainstream enzyme annotation routines, but they excel particularly when sequence identity between query and reference database falls below 25% [2] [1].
Table 3: Key research reagents and computational tools for context-enhanced Sci-LLM research
| Tool/Resource | Type | Primary Function | Application in Research |
|---|---|---|---|
| ESM2 | Protein Language Model | Generating protein sequence embeddings | State-of-the-art for enzyme function prediction |
| ProtBERT | Protein Language Model | Protein sequence representation | Alternative PLM for comparative studies |
| UniProtKB | Database | Curated protein sequences and annotations | Primary data source for training and evaluation |
| BLASTp | Algorithm | Sequence similarity search | Traditional baseline for function prediction |
| Pfam | Database | Protein family collections | Source of functional domain context |
| PLMSearch | Search Tool | Remote homology detection | Combines PLM embeddings with efficient search |
Visualization of the Tokenization Dilemma and Solution
Figure 2: The tokenization dilemma in Sci-LLMs and its solution through contextual input.
Discussion: Implications for Biological Research and Drug Development
The superior performance of context-only approaches fundamentally repositions the role of Sci-LLMs in biological discovery. Rather than treating these models as universal sequence decoders, the evidence supports reframing them as powerful reasoning engines over structured, human-readable knowledge [46]. This paradigm shift acknowledges that the primary strength of existing Sci-LLMs lies not in interpreting biomolecular syntax from scratch, but in their profound capacity for synthesizing established biological knowledge.
This approach has particular significance for drug development pipelines, where accurate function prediction can identify novel therapeutic targets or repurpose existing proteins. The ability of context-enhanced models to maintain performance on low-identity sequences (<25%) makes them particularly valuable for studying poorly characterized proteins or engineered enzymes with limited evolutionary relationships [2]. Furthermore, tools like PLMSearch demonstrate that PLM embeddings can power sensitive homology detection that approaches structure-based methods in accuracy while maintaining the efficiency of sequence-based search [27].
The emerging architecture of hybrid scientific AI agents combines the pattern recognition strength of PLMs with the established reliability of bioinformatics tools like BLASTp. This synergistic approach leverages the complementary strengths of both methodologies—PLMs for difficult cases with limited homology and BLASTp for sequences with clear evolutionary relationships [2] [1]. As these integrations mature, they promise to significantly accelerate annotation workflows for large-scale genomic and metagenomic projects.
Overcoming the tokenization dilemma through high-level contextual input represents a critical advancement for scientific AI. The experimental evidence consistently demonstrates that providing structured biological context—rather than raw sequences—unlocks the sophisticated reasoning capabilities of Sci-LLMs while avoiding the informational noise inherent in low-level sequence data. This approach facilitates a new generation of hybrid scientific AI agents that reposition developmental focus from direct sequence interpretation toward high-level knowledge synthesis.
For researchers, scientists, and drug development professionals, these findings indicate that the most productive path forward involves leveraging established bioinformatics resources to create rich contextual representations that maximize Sci-LLM performance. As protein language models continue to evolve, their integration with traditional methods like BLASTp—each playing to their respective strengths—will likely become standard practice in biological discovery pipelines, ultimately accelerating our understanding of protein function and enabling novel therapeutic development.
In the field of bioinformatics, the accurate functional annotation of protein sequences is a cornerstone for advancements in genomics, systems biology, and drug development. For decades, homology-based search tools, particularly BLASTp, have been the undisputed gold standard for this task, operating on the principle that sequence similarity implies functional similarity. However, the recent emergence of protein language models (PLMs) like ESM and ProtBERT, pre-trained on millions of protein sequences, presents a powerful alternative. These models learn complex patterns and biophysical properties from sequence data alone, enabling them to predict function even in the absence of close homologs. This guide provides an objective comparison of these approaches, focusing on the critical trade-off between predictive accuracy and computational cost—a key consideration for research efficiency and scalability.
Performance Comparison: PLMs vs. BLASTp
Predictive Accuracy on Enzyme Commission Number Annotation
A comprehensive comparative assessment of PLMs for Enzyme Commission (EC) number prediction provides critical experimental data. The study evaluated models including ESM2, ESM1b, and ProtBERT against the traditional standard, BLASTp. The results, summarized in the table below, reveal a nuanced performance landscape [1] [2].
Table 1: Comparative Performance of BLASTp and Protein Language Models in EC Number Prediction
| Method | Overall Accuracy | Strength | Key Weakness |
|---|---|---|---|
| BLASTp | Marginally better overall [1] [2] | Excels at predicting certain EC numbers, particularly with clear homologs [1] | Performance drops sharply for sequences with low identity (<25%) to database entries [1] |
| PLMs (e.g., ESM2) | Highly competitive, surpasses one-hot encoding DL models [1] [2] | More accurate for difficult-to-annotate enzymes and those without homologs; superior below 25% sequence identity [1] | Still requires improvement to fully replace BLASTp in mainstream annotation [1] |
| Combined Approach | Surpasses individual method performance [1] | Complementary strengths provide more robust annotation [1] | Increased computational and pipeline complexity [1] |
The core finding is that while BLASTp maintains a slight overall edge, the best PLMs like ESM2 are superior in challenging scenarios, such as annotating enzymes with no close homologs or where sequence identity to known proteins falls below 25% [1]. This suggests that PLMs capture more fundamental functional signatures beyond simple sequence alignment.
Computational Resource Requirements
The resource footprint of these tools is a major factor in their practical application. The following table synthesizes performance and resource data from large-scale assessments and tool-specific studies [47] [8] [27].
Table 2: Computational Cost and Sensitivity Comparison of Search Methods
| Method | Type | Relative Speed | Sensitivity (Remote Homology) | Typical Use Case |
|---|---|---|---|---|
| BLASTp | Sequence Alignment | Baseline (Fast) [8] | Low to Moderate [27] | Routine annotation of sequences with clear homologs |
| MMseqs2 | Sequence Alignment | Faster than BLASTp [8] | Moderate (comparable to BLASTp) [8] [27] | Rapid large-scale database searches |
| HHblits | Profile HMM | Slower | High [27] | Detecting very distant relationships |
| PLMSearch | PLM-based | Millions of pairs in seconds (comparable to MMseqs2) [27] | High (3x MMseqs2 on superfamily-level; comparable to structure search) [27] | Sensitive, large-scale remote homology detection |
| pLM-BLAST | PLM-based | Significantly faster than HHsearch [47] | On par with HHsearch [47] | Detecting distant homology with local alignments |
| Structure Search (TM-align) | Structure Alignment | Extremely Slow (4 orders of magnitude slower than PLMSearch) [27] | Very High [27] | Gold standard when structures are available |
The key insight is that modern PLM-based search tools like PLMSearch and pLM-BLAST achieve a transformative balance, offering sensitivity that rivals or exceeds much slower profile- and structure-based methods, while operating at speeds comparable to fast sequence alignment tools [47] [27]. This makes them exceptionally suitable for projects requiring both high accuracy and high throughput.
Experimental Protocols and Workflows
Standardized Evaluation Methodology
To ensure fair comparisons, benchmark studies typically follow a structured protocol. For EC number prediction, the process is defined as a multi-label classification problem to account for promiscuous and multi-functional enzymes. The standard workflow involves [1]:
- Data Sourcing: Protein sequences and their validated EC numbers are extracted from curated databases like UniProtKB.
- Sequence Splitting: Sequences are partitioned into training and test sets. Crucially, the test set often includes sequences with low identity (<25%) to any sequence in the training set to evaluate performance on "difficult" annotations.
- Feature Extraction: For PLMs, the raw protein sequence is passed through a pre-trained model (e.g., ESM2) to generate a numerical representation, or "embedding," of the sequence.
- Model Training & Prediction: A classifier (e.g., a fully connected neural network) is trained on the embeddings from the training set to predict EC numbers. Its performance is then evaluated on the held-out test set.
- BLASTp Baseline: For the same test set, BLASTp searches are conducted against a database of training sequences, transferring EC numbers from the top hits based on sequence similarity.
This methodology allows for a direct, quantitative comparison of accuracy (e.g., precision, recall) between the two paradigms on identical test data [1].
Workflow Visualization: PLM-Based Annotation
The following diagram illustrates the typical workflow for protein function annotation using a protein language model, from sequence input to functional prediction.
The Scientist's Toolkit: Essential Research Reagents
Successful implementation of these annotation strategies relies on a suite of software tools and databases. The table below details key resources for building a modern protein function prediction pipeline.
Table 3: Essential Research Reagents for Protein Function Annotation
| Tool / Resource | Type | Primary Function | Reference |
|---|---|---|---|
| BLASTp | Sequence alignment tool | The standard for fast, homology-based function transfer. | [1] [8] |
| ESM2 | Protein Language Model | A state-of-the-art PLM for generating informative sequence embeddings for prediction. | [1] [48] |
| ProtT5 | Protein Language Model | Another powerful PLM, often used as the backbone for tools like pLM-BLAST. | [47] [27] |
| pLM-BLAST | PLM-based search tool | Detects distant homology by aligning sequences using context-aware, PLM-derived substitution scores. | [47] |
| PLMSearch | PLM-based search tool | Enables ultra-fast, sensitive homology search at scale by predicting structural similarity from embeddings. | [27] |
| METL | Biophysics-based PLM | A PLM pre-trained on biophysical simulation data, excelling in protein engineering tasks with small datasets. | [48] |
| UniProtKB/Swiss-Prot | Protein Database | A high-quality, manually annotated database used for training and benchmarking. | [1] |
Strategic Implementation and Future Outlook
Decision Framework for Practitioners
Choosing the right tool depends on the specific research goals and constraints. The following decision graph provides a practical guide for selecting an annotation strategy.
Emerging Trends and Complementary Strengths
The future of protein annotation lies not in a single superior tool, but in hybrid approaches that leverage the complementary strengths of each paradigm. As one study concludes, "BLASTp and LLM models complement each other and can be more effective when used together" [1]. Furthermore, new models are beginning to incorporate biophysical principles during pre-training. Frameworks like METL, which unites machine learning with biophysical modeling, show exceptional promise in protein engineering, especially for generalizing from very small experimental datasets [48]. As these models evolve and computational efficiency improves, PLMs are poised to become an integral, if not dominant, component of the bioinformatician's toolkit for protein function prediction.
Protein function annotation is a cornerstone of modern biology, underpinning discoveries in genomics, metabolic pathway engineering, and therapeutic development. For decades, sequence similarity search with tools like BLASTp has been the established gold standard for transferring functional knowledge from characterized proteins to novel sequences. However, the recent emergence of protein language models (pLMs)—deep learning models pre-trained on millions of protein sequences—offers a powerful, alignment-free approach to function prediction. Rather than existing in opposition, these methodologies possess complementary strengths. This guide objectively compares the performance of BLASTp and pLM-based annotation strategies and provides a framework for their integrated use. Synthesizing recent experimental evidence, we demonstrate that a hybrid workflow leveraging the reliability of BLASTp for clear homologs with the sensitivity of pLMs for remote homology and orphan proteins delivers superior robustness, especially for applications in drug discovery where accurate functional insights are critical.
Performance Comparison: BLASTp vs. Protein Language Models
A direct comparative assessment of BLASTp and pLMs for enzyme function prediction reveals a nuanced performance landscape. While BLASTp maintains a slight overall advantage, pLMs excel in specific, challenging scenarios [1].
Table 1: Overall Performance Comparison on Enzyme Commission (EC) Number Prediction
| Method | Overall Accuracy | Performance on High-Identity (>30%) Sequences | Performance on Low-Identity (<25%) Sequences | Key Strength |
|---|---|---|---|---|
| BLASTp | Marginally better [1] | Excellent | Poor | Reliability when clear homologs exist |
| pLMs (e.g., ESM2) | Slightly lower [1] | Good | Significantly better [1] | Predicting for remote homologs and orphan proteins |
The performance gap stems from their fundamental operating principles. BLASTp identifies homology via sequence alignment, which becomes unreliable when sequence identity drops below the "twilight zone" (~25%) [47]. In contrast, pLMs like ESM2 and ProtT5 generate sequence embeddings—dense numerical representations that encapsulate evolutionary, structural, and functional constraints learned from vast sequence datasets. This allows them to detect subtle, context-dependent patterns that elude alignment-based methods [9] [27].
Table 2: Technical and Operational Characteristics
| Characteristic | BLASTp | pLM-Based Search (e.g., PLMSearch, pLM-BLAST) |
|---|---|---|
| Core Principle | Local sequence alignment and k-mer filtering [8] | Comparison of contextual embeddings from deep learning models [27] [47] |
| Primary Input | Single protein sequence | Single protein sequence |
| Speed | Fast [27] | Comparable to fast sequence search tools like MMseqs2 [27] |
| Sensitivity to Remote Homology | Low | High, can recall proteins with dissimilar sequences but similar structures [27] |
| Dependence on Database Annotation | High | Lower, can make predictions from sequence patterns alone |
Experimental Protocols and Workflows
Standard BLASTp Protocol for Function Prediction
The following protocol is standard for homology-based function transfer using BLASTp and is employed in numerous function prediction pipelines [8].
- Query and Database Preparation: Formulate the query protein sequence. Select a target protein database with functional annotations (e.g., Swiss-Prot).
- Sequence Search: Execute BLASTp against the target database. Default parameters are often used, but performance can be tuned.
- Command example:
blastp -query my_protein.fasta -db swissprot -out results.xml -outfmt 5 -evalue 1e-3
- Command example:
- Hit Selection and Scoring: Parse the results to collect top homologous hits. The associated functions (e.g., Gene Ontology terms or EC numbers) of these hits are then transferred to the query.
- Function Transfer: Use a scoring function to derive a final prediction from the set of hits. A simple method is to transfer annotations from the single best hit (highest score or lowest E-value). Advanced scoring functions that aggregate evidence from multiple hits (e.g., the S1 function, which sums the negative log of the E-value of all hits sharing a particular annotation) have been shown to improve accuracy [8].
pLM-Based Function Prediction Workflow
pLMs can be used for function prediction in two primary ways: as a feature extractor for a downstream classifier or as the backbone for a dedicated search tool.
A. pLM as Feature Extractor for Classification
This is a common approach for predicting specific functional classes, such as DNA-binding proteins or secreted effectors [49] [50].
- Embedding Generation: Pass the protein sequence through a pre-trained pLM (e.g., ESM2, ProtT5) to obtain a per-residue or per-sequence embedding vector.
- Model Training: Use the embeddings as input features to train a supervised machine learning classifier (e.g., a fully connected neural network, support vector machine) on a labeled dataset.
- Prediction: The trained model predicts functional classes for new query sequences based on their pLM-generated embeddings.
B. pLM-Based Homology Search (e.g., PLMSearch, pLM-BLAST)
These tools use pLM embeddings to find functionally related proteins directly [27] [47].
- Embedding Generation: Generate embeddings for the query and all sequences in the target database.
- Similarity Calculation: Compute the pairwise similarity between the query embedding and database embeddings. PLMSearch, for instance, uses a neural network (SS-predictor) trained on structural similarity (TM-score) to predict the functional similarity of a protein pair [27].
- Alignment (Optional): Some tools, like pLM-BLAST and PLMAlign, can generate context-aware alignments by creating a dynamic, embedding-derived substitution matrix for residue-level comparison [27] [47].
- Ranking and Annotation: Rank the target proteins by predicted similarity and transfer annotations from the top hits.
Diagram 1: pLM-Based Function Prediction Workflow. This diagram outlines the two primary pathways for using protein language models in function prediction.
The Hybrid BLASTp-pLM Workflow: A Practical Guide
Integrating BLASTp and pLMs creates a system that is more robust than the sum of its parts. The following workflow leverages the strengths of each method.
- Initial Triage with BLASTp: Perform a BLASTp search against a comprehensive, annotated database.
- Result Evaluation:
- Confident Hit: If a high-identity, statistically significant hit (e.g., E-value < 1e-30, identity > 40%) is found, its annotation can be transferred with high confidence. This provides a quick and reliable answer for a large proportion of queries.
- Ambiguous/Weak Hit: If BLASTp returns only weak hits (low identity, high E-value) or no hits at all, proceed to the pLM arm.
- pLM-Based Analysis:
- Path A (Search): Use a tool like PLMSearch or pLM-BLAST to search for remote homologs that might have been missed. These tools can detect relationships based on structural and functional similarity concealed at the sequence level [27].
- Path B (Direct Prediction): For specific functions (e.g., "Is this a DNA-binding protein?"), use a dedicated pLM-based classifier (e.g., ESM-DBP [50]) to get a direct prediction.
- Synthesis and Annotation: Combine the evidence from both arms. A strong BLASTp hit provides a clear annotation. A weak BLASTp hit supported by a strong pLM-based prediction indicates potential remote homology, and the pLM result can be used for a more confident, nuanced annotation.
Diagram 2: Hybrid BLASTp-pLM Annotation Workflow. This integrated approach ensures robust results by leveraging the best capabilities of each method.
The Scientist's Toolkit: Essential Research Reagents and Solutions
Building and executing these workflows requires a suite of computational tools and databases. The following table details key resources for implementing a hybrid protein annotation strategy.
Table 3: Key Research Reagent Solutions for Protein Annotation
| Item Name | Type | Function in Workflow | Example/Reference |
|---|---|---|---|
| Annotated Protein Databases | Data Resource | Serves as the target for homology searches and the source of ground-truth annotations. | UniProt/Swiss-Prot [9], Pfam [27] |
| BLAST Suite | Software Tool | Performs fast, local sequence alignment for the initial homology search and annotation transfer. | BLASTp, PSI-BLAST [8] |
| Pre-trained pLMs | AI Model | Generates contextual embeddings from protein sequences for downstream prediction tasks. | ESM2 [50], ProtT5 [47], ProtBert [1] |
| pLM-Based Search Tools | Software Tool | Enables sensitive, embedding-based homology search to find remote homologs with similar structure/function. | PLMSearch [27], pLM-BLAST [47] |
| Specialized pLM Predictors | AI Model | Provides direct prediction of specific protein properties or functions from sequence embeddings. | ESM-DBP (DNA-binding) [50], T4SEpp (bacterial effectors) [49] |
The debate between traditional BLASTp and modern protein language models is not a zero-sum game. Experimental data confirms that BLASTp remains a highly reliable and efficient tool for annotating proteins with clear homologs, while pLMs provide a breakthrough in sensitivity for detecting remote evolutionary relationships and annotating proteins with few or no database homologs. For researchers and drug development professionals, the most robust and future-proof strategy is a hybrid one. By designing a workflow that uses BLASTp for initial, high-confidence triage and strategically deploys pLM-based tools to resolve ambiguous cases and uncover deep functional insights, scientists can achieve a level of annotation accuracy and coverage that neither method can provide alone. This synergistic approach will be critical for exploring the vast uncharted territories of protein sequence space in the era of precision biology and therapeutics.
Head-to-Head Validation: A Data-Driven Comparison of Accuracy, Scope, and Limitations
The accurate prediction of Enzyme Commission (EC) numbers is a fundamental challenge in genomics and bioinformatics, directly impacting applications in metabolic engineering, drug discovery, and functional genomics. For decades, homology-based methods like BLASTp have served as the gold standard for functional annotation. However, the recent emergence of protein Large Language Models (LLMs) offers a promising alternative that leverages learned representations of protein sequences rather than explicit sequence similarity.
This benchmark guide provides a comprehensive comparative assessment of these competing paradigms, offering experimental data and methodological insights to help researchers select appropriate tools for enzyme function prediction. We focus specifically on the performance comparison between protein LLMs and BLASTp, contextualizing results within the broader thesis that these approaches offer complementary strengths rather than mutually exclusive solutions.
Table 1: Overall Performance Comparison of EC Number Prediction Methods
| Method | Type | Overall Accuracy | Strengths | Limitations |
|---|---|---|---|---|
| BLASTp | Homology-based | Marginally better overall [10] [2] | Excellent for enzymes with clear homologs [10] | Limited for sequences without homologs; performance drops below 25% identity [10] |
| Protein LLMs (Ensemble) | Deep Learning | Slightly lower than BLASTp but complementary [10] [2] | Better for difficult-to-annotate enzymes; effective below 25% identity [10] | Requires computational resources; training data dependency |
| ESM2 | Protein LLM | Best among tested LLMs [10] [2] | Accurate predictions for enzymes without homologs [10] | - |
| ProtBERT | Protein LLM | Lower than ESM2 [10] | Can be fine-tuned for specific tasks [10] | Underperforms ESM2 in comparative assessment [10] |
| GraphEC | Structure-based Geometric Learning | Superior to sequence-based methods [51] | Incorporates structural information and active sites [51] | Depends on predicted structures; computationally intensive |
Experimental evidence from a comprehensive 2025 study reveals that while BLASTp provides marginally better overall results, the performance difference is minimal, and protein LLMs excel in specific challenging scenarios [10] [2]. This suggests that the selection between these approaches should be guided by specific use cases rather than absolute performance rankings.
The complementary nature of these methods is particularly noteworthy. The same study found that "LLMs better predict certain EC numbers while BLASTp excels in predicting others," indicating that ensemble approaches incorporating both methodologies might offer optimal performance [10].
Methodology and Experimental Protocols
Experimental Design for Comparative Assessment
Table 2: Key Methodological Components in EC Prediction Benchmarking
| Component | Description | Implementation in Cited Studies |
|---|---|---|
| Data Source | UniProtKB SwissProt and TrEMBL | Downloaded February 2023; only UniRef90 cluster representatives retained to avoid redundancy [10] |
| Problem Formulation | Multi-label classification | Incorporates promiscuous and multi-functional enzymes; all hierarchical levels included [10] |
| Feature Extraction | Protein sequence embeddings | ESM2, ESM1b, ProtBERT embeddings used as input to fully connected neural networks [10] |
| Comparison Baseline | Traditional alignment | BLASTp used as reference standard for performance comparison [10] [2] |
| Evaluation Framework | Multiple test scenarios | Standard benchmarks plus challenging cases (low identity, no homologs) [10] |
The experimental protocol for benchmarking EC number prediction methods requires careful design to ensure fair comparison. The most comprehensive studies define EC number prediction as a multi-label classification problem incorporating promiscuous and multi-functional enzymes [10]. Each protein sequence (xi) has an associated binary label vector (yi) where elements indicate association with particular EC numbers.
Data processing typically involves using UniRef90 cluster representatives from UniProtKB to enhance biologically relevant information retrieval and avoid overrepresentation of similar sequences [10]. This procedure ensures that enzymes sharing more than 90% identity are removed, creating a more challenging and realistic evaluation scenario.
Protein LLM Architectures and Training
Protein LLMs like ESM2, ESM1b, and ProtBERT are transformer-based networks pre-trained on massive protein sequence databases. For EC number prediction, these models are typically used as feature extractors, where outputs from specific layers serve as embeddings input to downstream classifiers [10].
In benchmark studies, these embeddings are processed by fully connected neural networks that learn to map the extracted features to EC number predictions. The ESM2 model specifically stood out as the best performer among tested LLMs, providing more accurate predictions on difficult annotation tasks [10] [2].
Advanced Structural and Geometric Learning Approaches
Beyond sequence-based methods, recent advancements incorporate protein structural information through geometric graph learning. GraphEC represents this next-generation approach, utilizing ESMFold-predicted structures and enzyme active sites to enhance prediction accuracy [51].
The GraphEC methodology involves several sophisticated components:
Active Site Prediction: A dedicated module (GraphEC-AS) first identifies enzyme active sites, achieving an AUC of 0.9583 on independent tests [51]
Structure-Based Graph Construction: Protein structures predicted by ESMFold are used to construct protein graphs capturing spatial relationships
Geometric Graph Learning: Graph neural networks process these structures to learn representations that incorporate three-dimensional constraints
Label Diffusion: Homology information is incorporated through diffusion algorithms to further refine predictions [51]
This approach demonstrates the evolving landscape of EC number prediction, where combining sequential, structural, and evolutionary information delivers superior performance.
Table 3: Key Research Resources for EC Number Prediction Studies
| Resource | Type | Application in EC Prediction | Access |
|---|---|---|---|
| UniProtKB | Database | Primary source of protein sequences and EC annotations [10] | https://www.uniprot.org/ |
| ESM2/ESM1b | Protein LLM | Generate sequence embeddings for deep learning models [10] | https://github.com/facebookresearch/esm |
| ProtBERT | Protein LLM | Alternative protein language model for comparative studies [10] | https://huggingface.co/Rostlab/prot_bert |
| BLASTp | Algorithm | Gold standard homology-based method for performance comparison [10] [2] | https://blast.ncbi.nlm.nih.gov/ |
| GraphEC | Software | Geometric graph learning for structure-informed predictions [51] | Publication-based |
| CLEAN | Software | Contrastive learning-based EC number predictor [51] | Publication-based |
| AlphaFold2/ESMFold | Software | Protein structure prediction for structural approaches [51] | https://alphafold.ebi.ac.uk/ |
The experimental workflows described require access to both data resources and computational tools. For protein sequences and ground truth EC numbers, UniProtKB serves as the authoritative source, with careful filtering to use only UniRef90 cluster representatives to avoid homology bias [10].
For implementing protein LLM approaches, the ESM model series provides the best performance according to comparative studies, while ProtBERT offers an alternative implementation based on BERT architecture [10]. For structural approaches, ESMFold enables rapid structure prediction with accuracy comparable to AlphaFold2 but with significantly reduced computational requirements [51].
Interpretation of Experimental Results
Performance Across Different Annotation Scenarios
The comparative performance between BLASTp and protein LLMs varies significantly across different annotation scenarios:
High-Identity Cases: For enzymes with clear homologs in databases (sequence identity >25%), BLASTp remains the superior choice, leveraging direct evolutionary relationships for accurate function transfer [10].
Low-Identity Cases: For sequences with limited homology (identity <25%), protein LLMs significantly outperform BLASTp, demonstrating their ability to capture functional signatures beyond sequence similarity [10].
Novel Enzyme Families: For enzymes without close homologs, ESM2 emerged as the best model among tested LLMs, providing more accurate predictions where traditional methods fail [10] [2].
These findings support a hybrid approach to enzyme annotation where the method selection is guided by the characteristics of the target sequence and the availability of homologous sequences in databases.
Complementary Strengths and Hybrid Approaches
The research consistently demonstrates that BLASTp and protein LLMs exhibit complementary prediction profiles, with each method excelling for different subsets of EC numbers [10]. This complementarity suggests that combined approaches may achieve performance exceeding either method individually.
Evidence from independent tests shows that geometric learning methods like GraphEC achieve superior performance compared to sequence-based methods, with the additional advantage of providing active site predictions and optimum pH estimates [51]. This represents a significant advancement in comprehensive enzyme function annotation.
Based on the comprehensive performance benchmarking, we recommend the following guidelines for researchers selecting EC number prediction methods:
For routine annotation of enzymes with expected homologs, BLASTp remains the most reliable and efficient choice [10] [2]
For challenging cases involving sequences with low identity to characterized enzymes, protein LLMs (particularly ESM2) should be employed to leverage learned functional representations [10]
For maximum accuracy and additional functional insights (active sites, optimum pH), structure-aware methods like GraphEC represent the cutting edge, though with increased computational requirements [51]
For critical applications, implement a hybrid approach that combines multiple methods to leverage their complementary strengths
The field of EC number prediction continues to evolve rapidly, with protein language models increasingly closing the performance gap with traditional methods while offering advantages in specific challenging scenarios. Researchers should consider maintaining both approaches in their bioinformatics toolkit to address the diverse range of annotation challenges encountered in modern genomic research.
Protein function annotation is a cornerstone of modern bioinformatics, critical for advancing research in areas ranging from metabolic engineering to drug discovery. For decades, sequence alignment tools like BLASTp have served as the gold standard for transferring functional knowledge from characterized proteins to novel sequences based on similarity [1] [3]. However, the recent emergence of protein language models (pLMs) represents a paradigm shift in how we extract functional information from amino acid sequences alone. These models, inspired by breakthroughs in natural language processing, learn evolutionary patterns and biochemical principles from millions of protein sequences through self-supervised training, enabling them to predict function without explicit reliance on sequence homology [3] [14] [52].
The fundamental distinction between these approaches lies in their underlying mechanisms. BLASTp and other alignment-based tools operate on the principle of homology transfer, identifying statistically significant local alignments between a query sequence and databases of annotated proteins [53]. In contrast, pLMs function as pattern recognition systems, leveraging knowledge embedded in their parameters during pre-training to infer function directly from sequence composition and context [14] [52]. This analytical difference translates to complementary strengths and weaknesses that become apparent across different annotation scenarios.
This guide provides an objective comparison of these tools through the lens of recent research, with a particular focus on Enzyme Commission (EC) number prediction—a challenging multi-label classification problem that serves as an excellent benchmark for functional annotation capabilities [1]. By examining experimental data and performance metrics across diverse conditions, we aim to equip researchers with practical insights for selecting the optimal tool based on their specific annotation context.
Performance Comparison: Quantitative Analysis Across Experimental Conditions
Comparative studies reveal that while BLASTp maintains a slight overall advantage in standard annotation scenarios, pLMs have demonstrated remarkable capabilities that complement traditional approaches. A comprehensive 2025 assessment of protein language models for EC number prediction found that BLASTp provided marginally better results overall when evaluated against standard benchmarking datasets [1] [2]. However, this performance advantage was not uniform across all enzyme classes or annotation contexts.
The same study demonstrated that deep learning models using pLM embeddings significantly outperformed models relying on one-hot encodings of amino acid sequences, highlighting the superior representational capacity of learned protein embeddings [1]. Among the pLMs evaluated, the ESM2 model emerged as the best performer, providing more accurate predictions on difficult annotation tasks and for enzymes without close homologs in reference databases [1] [2].
Table 1: Overall Performance Comparison for EC Number Prediction
| Tool | Overall Accuracy | Strength Scenarios | Key Limitations |
|---|---|---|---|
| BLASTp | Slightly higher overall [1] | High-identity homologs present [1] | Fails with no homologs [1] |
| pLMs (ESM2) | Competitive, complementing BLASTp [1] | Low-identity sequences (<25%), difficult annotations [1] [2] | Still developing for mainstream annotation [1] |
| Combined Approach | Surpasses individual methods [1] | Comprehensive annotation across diverse proteins [1] | Increased computational complexity [1] |
Performance Across Sequence Identity Ranges
The most significant performance differentiation emerges when analyzing performance across varying levels of sequence identity. Research demonstrates that pLMs excel precisely where BLASTp struggles—when annotating sequences with low similarity to characterized proteins in databases.
A critical finding from recent comparative studies indicates that pLMs provide good predictions for more difficult-to-annotate enzymes, particularly when the identity between the query sequence and the reference database falls below 25% [1] [2]. This represents a crucial threshold where traditional homology-based methods experience significant performance degradation, while pLMs maintain robust predictive capability by leveraging learned evolutionary and structural patterns beyond simple sequence similarity.
Table 2: Performance Across Sequence Identity Ranges
| Sequence Identity | BLASTp Performance | pLM Performance | Recommended Tool |
|---|---|---|---|
| >40% | Excellent [1] | Good [1] | BLASTp |
| 25-40% | Good [1] | Competitive [1] | Both (Complementary) |
| <25% | Limited [1] [2] | Good, superior for difficult cases [1] [2] | pLMs (ESM2) |
| No Homologs | Fails [1] | Still functional [1] | pLMs |
Performance on Specific Protein Categories
The relative performance of these tools further varies across different protein functional categories and taxonomic groups. Studies have revealed that current pLMs often exhibit biases against proteins from underrepresented species, with viral proteins being particularly affected [26]. These proteins, frequently described as the "dark matter" of the biological world due to their vast diversity and sparse representation in training datasets, present particular challenges for pLMs trained primarily on standard UniProtKB datasets [26].
However, research also shows that fine-tuning pre-trained pLMs on domain-specific datasets can mitigate these biases by refining embeddings to capture diverse sequences and context-specific features [26]. For specialized applications such as antibody engineering, antibody-specific language models (AbLMs) have been developed that demonstrate superior performance for tasks like paratope prediction and developability optimization [52].
Experimental Protocols and Methodologies
Standardized Evaluation Framework for EC Number Prediction
To enable fair comparison between BLASTp and pLMs, researchers have developed standardized evaluation protocols focusing on EC number prediction as a benchmark task. The core methodology involves multi-label classification incorporating promiscuous and multi-functional enzymes (with more than one EC number) [1].
In a typical experimental setup, let ( X ) be the set of protein sequences and ( Y ) be the set of EC numbers. Each protein sequence ( xi ) in ( X ) has an associated binary label vector ( yi ) of length ( |Y| ), where ( |Y| ) is the total number of unique EC numbers. Each vector element ( y{ij} ) is either 0 or 1, indicating whether protein sequence ( xi ) is associated with the EC number ( j ) in ( Y ) [1]. The classification follows a global approach for hierarchical multi-label classification, where a single classifier predicts the entire hierarchy of labels and their relationships [1].
Dataset construction typically utilizes UniProtKB data (SwissProt and TrEMBL) with only UniRef90 cluster representatives retained to enhance dataset quality [1]. This careful curation ensures that performance evaluations reflect real-world annotation challenges rather than dataset-specific artifacts.
pLM Fine-Tuning Protocols for Enhanced Performance
For protein language models, specialized fine-tuning techniques have been developed to optimize performance for specific annotation tasks. Parameter-efficient fine-tuning (PEFT) strategies, particularly Low-Rank Adaptation (LoRA), have emerged as effective approaches for adapting large pLMs to specialized domains [26].
The LoRA method decomposes model weight matrices into smaller, low-rank matrices, dramatically reducing the number of trainable parameters and computational requirements [26]. This approach allows for rapid adaptation without additional inference latency, making it feasible to fine-tune even massive models like ESM2-3B on domain-specific data. A rank of 8 has been shown to achieve competitive performance while maintaining computational efficiency [26].
Research has demonstrated that fine-tuning with diverse learning frameworks—including masked language modeling, classification, and contrastive learning—significantly enhances embedding quality for specialized protein families, such as viral proteins that are typically underrepresented in general training datasets [26].
BLASTp Configuration for Optimal Annotation
For BLASTp evaluations, standard configurations typically employ an E-value threshold of 0.001 for significance filtering, with database searches conducted against comprehensively annotated reference datasets [1]. The highest-scoring hit meeting significance thresholds is typically used for function transfer, though more sophisticated consensus approaches can be employed for challenging cases.
Advanced implementations often combine BLASTp with machine learning models to assign EC numbers from homologous enzymes, compensating for shortcomings in simple homology-based approaches [1]. This hybrid methodology recognizes that while sequence similarity provides strong evidence of functional relationship, it cannot capture all nuances of enzyme function, particularly for promiscuous enzymes or those with divergent evolutionary histories.
Decision Framework: When to Use Each Tool
Tool Selection Based on Annotation Context
The experimental data supports a contextual framework for tool selection based on specific annotation scenarios and research objectives. The following decision matrix synthesizes findings from multiple comparative studies to guide researchers in selecting the optimal tool for their specific use case.
Table 3: Situational Tool Selection Guide
| Annotation Scenario | Recommended Tool | Rationale | Evidence |
|---|---|---|---|
| Routine annotation with expected homologs | BLASTp | Superior performance when high-identity matches exist | [1] |
| Novel enzyme families, low-identity sequences | pLMs (ESM2) | Better prediction for sequences with <25% identity to database | [1] [2] |
| Underrepresented protein families | Fine-tuned pLMs | Domain adaptation captures specific features | [26] |
| Comprehensive annotation pipeline | Combined approach | Complementary strengths maximize coverage | [1] |
| Antibody-specific annotation | Specialized AbLMs | Optimized for structural uniqueness of antibodies | [52] |
Complementary Strengths and Hybrid Approaches
Rather than viewing BLASTp and pLMs as mutually exclusive alternatives, the experimental evidence strongly supports their complementary relationship in comprehensive annotation pipelines [1]. Studies have consistently found that LLMs better predict certain EC numbers while BLASTp excels in predicting others, suggesting that a hybrid approach can leverage the distinctive capabilities of each methodology [1].
This complementarity arises from the fundamental differences in how these tools extract information from protein sequences. BLASTp relies on explicit evolutionary relationships manifested through sequence conservation, while pLMs leverage implicit patterns learned from the entire evolutionary landscape represented in their training data. The combination of these orthogonal information sources creates a more robust annotation system than either approach alone.
Essential Research Reagents and Computational Tools
Core Tools for Protein Function Annotation
Table 4: Essential Research Reagents and Computational Tools
| Tool/Resource | Type | Primary Function | Application Context |
|---|---|---|---|
| ESM2 | Protein Language Model | Protein sequence embedding | State-of-the-art EC prediction [1] [2] |
| BLASTp | Sequence Alignment | Homology-based search | Gold standard for similar sequences [1] |
| UniProtKB | Database | Curated protein sequences | Reference for annotation transfer [1] |
| CARD | Database | Antibiotic resistance genes | Specialized AMR annotation [18] |
| LoRA | Fine-tuning method | Parameter-efficient adaptation | Domain-specific pLM tuning [26] |
| AntiBERTa | Specialized pLM | Antibody-specific prediction | Paratope prediction [52] |
The comparative analysis of BLASTp and protein language models reveals a nuanced landscape where each tool demonstrates distinct advantages depending on the annotation context. BLASTp remains the gold standard for routine annotation tasks where sequences have clear homologs in reference databases, offering slightly superior overall performance in these scenarios [1]. However, protein language models excel in precisely the areas where BLASTp is weakest—annotating sequences with low similarity to characterized proteins, predicting functions for difficult-to-annotate enzymes, and handling cases where no close homologs exist [1] [2].
The most promising path forward lies in hybrid approaches that leverage the complementary strengths of both methodologies [1]. As protein language models continue to evolve and address current limitations—including biases against underrepresented protein families and computational demands—they are poised to become increasingly integral to mainstream annotation workflows [26]. For now, researchers can optimize their annotation pipelines by applying the situational framework presented here, selecting tools based on sequence characteristics, taxonomic context, and functional categories to maximize annotation accuracy and coverage.
Future developments in model architectures, training methodologies, and integration with structural information will further blur the distinctions between these approaches, ultimately leading to more accurate, comprehensive, and efficient protein function annotation systems that advance our understanding of biological systems and accelerate therapeutic development.
The accurate functional annotation of enzymes, typically represented by Enzyme Commission (EC) numbers, is a cornerstone of modern bioinformatics, with profound implications for understanding cellular metabolism, designing novel metabolic pathways, and advancing drug discovery [1]. For decades, sequence alignment tools like BLASTp have served as the gold standard for this task, operating on the principle that sequence similarity implies functional similarity [1]. However, the recent emergence of protein Language Models (pLMs)—deep learning models pre-trained on millions of protein sequences—offers a powerful, alignment-free alternative for function prediction [9].
A common framing positions these methods as competitors, yet a growing body of evidence suggests their relationship is fundamentally synergistic. This guide synthesizes recent comparative research to objectively demonstrate that BLASTp and pLMs are not mutually exclusive but are, in fact, highly complementary technologies. By examining their performance across different annotation scenarios, we provide a data-driven framework for researchers to strategically combine these tools, thereby achieving more accurate, robust, and comprehensive functional annotations than either method could provide alone.
Performance Comparison: Quantitative Data
Direct comparisons reveal that while BLASTp holds a slight overall advantage, pLMs excel in specific, critical areas, particularly when sequence similarity is low.
Table 1: Overall Performance Comparison between BLASTp and pLMs on EC Number Prediction
| Method | Overall Accuracy | Strength in High-Similarity Scenarios | Performance on Sequences with Low Homology | Key Differentiating Factor |
|---|---|---|---|---|
| BLASTp | Marginally better overall [1] | Excellent; the established gold standard [1] | Rapidly declining performance below 25% identity [1] | Relies on existence of closely related sequences in database |
| pLMs (e.g., ESM2) | Slightly lower but highly competitive overall [1] | Can be outperformed by BLASTp when strong homologs exist [1] | Superior; provides good predictions for difficult-to-annotate enzymes [1] | Leverages learned biochemical principles; does not require a homolog |
Table 2: Performance of Specific Protein Language Models
| pLM Model | Key Characteristics | Relative Performance |
|---|---|---|
| ESM2 | Transformer-based; trained on UniProtKB data [1] | Stands out as the best model among tested LLMs [1] |
| ESM1b | Earlier version of ESM models [1] | Competitive, but generally outperformed by ESM2 [1] |
| ProtBERT | Transformer-based; pre-trained on UniRef and BFD databases [1] | Effective, but a comprehensive comparison showed ESM2's superiority [1] |
The critical finding is that the strengths of each method are non-overlapping. One study concluded that "LLMs better predict certain EC numbers while BLASTp excels in predicting others," and that their combination is more effective than either tool individually [1]. This complementarity is most evident along the axis of sequence similarity, as visualized below.
Experimental Protocols for Key Studies
Understanding the experimental design behind these conclusions is crucial for interpreting the data and applying the findings.
Comparative Assessment of pLMs for EC Number Prediction
This seminal study provided a direct, large-scale comparison of BLASTp and several pLMs [1].
- Objective: To assess the performance of ESM2, ESM1b, and ProtBERT in predicting EC numbers and compare them against BLASTp and models using one-hot encodings [1].
- Data Preparation: Protein sequences and their EC numbers were extracted from the SwissProt and TrEMBL sections of UniProtKB. To ensure data quality and avoid redundancy, only representative sequences from UniRef90 clusters were used. The EC number prediction was framed as a multi-label classification problem [1].
- Model Training & Evaluation: pLMs were used as feature extractors. The embeddings (vector representations) they generated were fed into fully connected neural networks for EC number classification. The performance of these DL models was then rigorously benchmarked against the results from a standard BLASTp search [1].
- Key Metric for Complementarity: Performance was analyzed across different levels of sequence identity between the query and its closest homolog in the database, highlighting the strengths of each method [1].
The PhiGnet Methodology for Residue-Level Function Prediction
While not a direct BLASTp comparison, the PhiGnet approach exemplifies the advanced, interpretable functionality enabled by pLMs.
- Objective: Develop a method (PhiGnet) that uses statistics-informed graph networks to predict protein function and identify functional sites at the residue level from sequence alone [54].
- Data Integration: For a given protein sequence, an embedding from the pre-trained ESM-1b model is derived. This embedding is used as graph nodes, which are connected by edges representing evolutionary couplings (EVCs) and residue communities (RCs) [54].
- Model Architecture: The model employs a dual-channel architecture with stacked graph convolutional networks (GCNs) to process the EVC and RC data. This is followed by fully connected layers to produce probability scores for functional annotations [54].
- Interpretation: The model uses gradient-weighted class activation maps (Grad-CAM) to compute an activation score for each residue, quantifying its importance for a specific protein function and allowing for the identification of functional sites like binding pockets [54].
The Scientist's Toolkit: Essential Research Reagents
Successfully leveraging BLASTp and pLMs requires a suite of key databases, software tools, and computational resources.
Table 3: Key Research Reagents for Functional Annotation
| Tool / Resource | Type | Primary Function in Annotation | Relevance |
|---|---|---|---|
| UniProt Knowledgebase (UniProtKB) | Database | Central repository of protein sequence and functional information [1] | Source of ground-truth data for training pLMs and as a reference database for BLASTp searches. |
| ESM2 Model | Protein Language Model | Pre-trained deep learning model that generates informative embeddings from protein sequences [1] | A state-of-the-art pLM for enzyme function prediction; can be used as a feature extractor for downstream models. |
| BLASTp Suite | Software Suite | Performs local alignment searches of a query protein against a protein database [1] | The benchmark homology-based method for functional annotation; essential for comparative and complementary workflows. |
| PhiGnet | Specialized Prediction Tool | Statistics-informed graph network for function prediction and residue-level significance assessment [54] | Represents the next generation of interpretable pLM-based tools that can pinpoint functional sites. |
The evidence clearly demonstrates that the dichotomy between BLASTp and protein Language Models is a false one. BLASTp remains the tool of choice for annotating proteins with clear and close homologs in existing databases, a scenario where its performance is unmatched. Conversely, pLMs have emerged as a powerful technology for the "dark matter" of proteomics—proteins with low sequence similarity to characterized families, difficult-to-annotate enzymes, and for tasks requiring residue-level functional insight [1] [54].
For researchers and drug development professionals, the strategic path forward is integration, not substitution. A recommended workflow begins with a BLASTp search. For queries with high-identity hits, the annotation can be assigned with high confidence. For queries with weak or no hits, pLMs should be deployed to generate functional hypotheses. Finally, for critical applications like drug target validation, using both methods in concert provides the most robust functional evidence, leveraging the strengths of both homology-based and principle-based prediction paradigms. This synergistic approach will be essential for illuminating the vast landscape of uncharacterized proteins and accelerating biomedical discovery.
The accurate functional annotation of enzymes is a cornerstone of genomics, metabolic engineering, and drug discovery. For decades, sequence alignment tools like BLASTp have been the gold standard for transferring functional knowledge, such as Enzyme Commission (EC) numbers, from characterized enzymes to new sequences based on homology [1]. This approach, however, fails for the vast number of enzymes that lack closely related sequences in annotated databases. The emergence of protein Language Models (pLMs) offers a paradigm shift. These models, pre-trained on millions of protein sequences, learn fundamental principles of protein function and structure, potentially enabling them to annotate enzymes independently of sequence similarity [1] [3].
This guide provides an objective comparison of pLM and BLASTp performance, with a specific focus on the critical challenge of annotating enzymes without close homologs. Framed within the broader thesis of pLM versus BLASTp annotation research, we synthesize recent experimental data to delineate the strengths and limitations of each approach, providing researchers with a clear understanding of the current technological landscape.
Performance Comparison: pLMs vs. BLASTp
Direct comparative studies reveal that while BLASTp maintains a slight overall advantage, pLMs excel in specific, biologically critical scenarios, particularly when sequence identity is low.
Table 1: Overall Performance Summary of pLMs vs. BLASTp
| Method | Overall Performance | Strength | Weakness |
|---|---|---|---|
| BLASTp | Marginally better overall accuracy [1] | Superior for sequences with high similarity to annotated database entries [1] | Cannot assign function to proteins without homologous sequences [1] |
| pLMs (e.g., ESM2) | Highly competitive, complementary to BLASTp [1] | Better predictions for difficult-to-annotate enzymes and sequences with <25% identity to database entries [1] | Still requires improvement to become the new gold standard for mainstream annotation [1] |
A comprehensive assessment of pLMs (ESM2, ESM1b, ProtBERT) against BLASTp demonstrated that their predictive capabilities are not identical but complementary. The study found that "LLMs better predict certain EC numbers while BLASTp excels in predicting others" [1]. This suggests that the two methods capture different aspects of the information required for accurate function prediction.
Table 2: pLM Performance on Enzymes with Low Sequence Similarity
| Sequence Identity to Database | BLASTp Performance | pLM Performance | Key Findings |
|---|---|---|---|
| >50% Identity | High accuracy, par with HHsearch [47] | High accuracy [47] | Homology readily detected by both sequence-based and pLM-based methods. |
| <30% Identity | Performance declines; relies on slower profile HMMs [47] | Maintains high accuracy; ESM2 provides more accurate predictions [1] [47] | pLMs show significant advantage in distant homology detection. |
| <25% Identity (Difficult-to-annotate) | Fails or provides poor predictions [1] | Excels; provides good predictions where BLASTp struggles [1] | pLMs unlock functional insights for enzymes with no close homologs. |
The standout advantage for pLMs is their performance on sequences that share low identity with known proteins. The ESM2 model, in particular, "stood out as the best model among the LLMs tested, providing more accurate predictions on difficult annotation tasks and for enzymes without homologs" [1]. This capability is crucial for expanding the annotation of the "dark matter" of the protein universe, including proteins from underrepresented viral species [26].
Experimental Protocols for Critical Benchmarks
The performance data cited in this guide are derived from rigorous, large-scale experimental benchmarks. The following outlines the core methodologies employed in these foundational studies.
Protocol 1: Large-Scale EC Number Prediction Comparison
This protocol underpins the direct comparison between pLMs and BLASTp as summarized in [1].
- Problem Formulation: EC number prediction was treated as a multi-label classification problem, accounting for promiscuous and multi-functional enzymes. The entire EC hierarchy was predicted simultaneously using a global approach for hierarchical multi-label classification [1].
- Data Curation: Data was extracted from UniProtKB (SwissProt and TrEMBL). To ensure sequence diversity and avoid bias, only representative sequences from UniRef90 clusters were used. This process enhances the quality of the dataset by selecting sequences based on annotation score and sequence length [1].
- Model Training & Evaluation:
- pLMs: Representations (embeddings) were extracted from pre-trained models including ESM2, ESM1b, and ProtBERT. These embeddings were used as input features for a fully connected neural network classifier [1].
- Baselines: Deep learning models using one-hot encodings (e.g., DeepEC, D-SPACE) were implemented for comparison. Performance of all DL models was benchmarked against BLASTp [1].
- Assessment: Models were evaluated on their ability to correctly predict EC numbers, with a specific analysis of performance based on the sequence identity between the query and known database sequences [1].
Protocol 2: Distant Homology Detection with pLM-BLAST
This protocol from [47] details an alternative, alignment-based use of pLM embeddings for homology detection, a related task to function prediction.
- Embedding Generation: Protein sequences are passed through a pLM (ProtT5) to generate a per-residue embedding matrix. Each residue's embedding vector is normalized using its Euclidean norm [47].
- Substitution Matrix Calculation: A context-aware substitution matrix for two sequences is computed as the cosine similarity of every pair of residue embeddings from the two sequences. This replaces static matrices like BLOSUM62 [47].
- Alignment Algorithm: A modified Smith-Waterman algorithm is used with the dynamic embedding-based substitution matrix to generate a scoring matrix. Gap penalties are typically set to zero, as dissimilar regions naturally score low [47].
- Traceback and Alignment Extraction: The traceback procedure starts from all sequence boundaries (not just the highest score) to identify multiple candidate alignment regions. High-scoring local alignments are identified based on a threshold applied to the moving average of the path score [47].
The Scientist's Toolkit: Essential Research Reagents
The following table details key computational tools and resources essential for conducting research in pLM-based enzyme function prediction.
Table 3: Key Research Reagents for pLM-based Enzyme Annotation
| Tool / Resource | Type | Primary Function | Relevance to Difficult Cases |
|---|---|---|---|
| ESM2 [1] [16] | Protein Language Model | Generates contextual embeddings from single sequences. | Top-performing model for annotating enzymes without close homologs [1]. |
| UniRef90 [1] | Protein Sequence Database | Curated dataset of non-redundant sequences. | Provides high-quality, diverse data for training and benchmarking models [1]. |
| pLM-BLAST [47] | Homology Search Tool | Detects distant homology using pLM embeddings. | Connects highly divergent proteins, uncovering previously unknown homologous relationships [47]. |
| PLM-interact [16] | Fine-tuned pLM | Predicts protein-protein interactions from sequence. | Demonstrates the potential of fine-tuning pLMs for specific relational tasks beyond single-protein function [16]. |
| LoRA (Low-Rank Adaptation) [26] | Fine-tuning Method | Efficiently adapts large pLMs to specific domains. | Mitigates pLM bias against underrepresented proteins (e.g., viral enzymes) [26]. |
Visualizing the Broader pLM vs. BLASTp Research Workflow
The relationship between different methodologies and their application to enzyme annotation can be visualized as a decision workflow. This diagram integrates concepts from the provided search results, illustrating the complementary roles of traditional and modern approaches.
The analysis of difficult cases confirms that protein Language Models have emerged as a powerful technology capable of annotating enzymes without close homologs, a task at which traditional BLASTp fails. While BLASTp remains a robust and marginally superior tool for annotating sequences with clear homology, pLMs like ESM2 unlock new possibilities for functional inference in the low-similarity regime. The prevailing thesis in the field is not of outright replacement, but of powerful complementarity. As [1] concludes, "BLASTp and LLM models complement each other and can be more effective when used together." Future advancements will likely stem from hybrid approaches, efficient fine-tuning on specific protein families [26], and the development of next-generation pLMs explicitly designed to illuminate biology's darkest corners.
In the field of computational biology, the ability to accurately predict protein function is a cornerstone for advancements in drug discovery, metabolic engineering, and fundamental biological research. For decades, sequence similarity tools like BLASTp have served as the gold standard, operating on the principle that proteins with similar sequences perform similar functions. However, the recent emergence of Protein Large Language Models (LLMs) like ESM2, ESM1b, and ProtBERT has introduced a paradigm shift. These models do not merely compare sequences; they learn the complex, contextual "language" of proteins from millions of sequences, building an internal, high-level understanding of biochemical principles. This guide objectively compares the performance of these context-based Protein LLMs against traditional sequence-similarity methods, providing researchers with the experimental data and methodologies needed to evaluate their applications.
Defining the Paradigms: Sequence Alignment vs. Contextual Understanding
The core distinction between the two approaches lies in their fundamental operating principles.
Sequence-Based Methods (BLASTp): This is a local alignment search tool. It identifies regions of local similarity between a query protein sequence and sequences in a database by calculating an alignment score based on substitution matrices. Its prediction is fundamentally based on evolutionary homology; it transfers functional annotation from the most similar sequence(s) found in a curated database. Its performance is therefore constrained by the completeness and quality of the database and struggles with proteins that have no close homologs [1].
Context-Based Methods (Protein LLMs): Models like ESM2 are transformer-based neural networks pre-trained on millions of protein sequences from databases like UniProtKB in a self-supervised manner. During pre-training, they learn to predict masked amino acids in a sequence, forcing them to develop a deep, contextual understanding of protein syntax and semantics—the biophysical properties and evolutionary constraints that shape sequences. For function prediction, these "learned" representations (embeddings) are then used as features to train a simple classifier, such as a fully connected neural network, to predict EC numbers [1] [3]. This allows them to infer function from patterns that may not depend on direct sequence homology.
Performance Comparison: Quantitative Benchmarks
A comprehensive comparative assessment reveals the distinct strengths and weaknesses of each paradigm. The following tables summarize key performance metrics from recent, large-scale evaluations.
| Method | Model / Tool Name | Overall Accuracy (Approx.) | Key Strengths | Key Limitations |
|---|---|---|---|---|
| Sequence-Based | BLASTp | Slightly Higher [1] | Excellent when strong homologs exist; well-understood and fast. | Cannot annotate proteins without homologs; performance drops sharply with sequence divergence. |
| Context-Based | ESM2 + DNN | High (Best among LLMs) [1] | Predicts functions without homologs; excels on difficult annotations and sequences with <25% identity to known proteins. | Marginally lower overall accuracy than BLASTp in standard benchmarks; requires significant computational resources for training. |
| ESM1b + DNN | High [1] | Strong performance, a established predecessor to ESM2. | Generally outperformed by the newer ESM2 architecture. | |
| ProtBERT + DNN | High [1] | Competitive performance, based on the BERT architecture. | Performance often trails ESM models in direct comparisons [1]. |
Table 2: Performance on Specific Challenging Scenarios
| Scenario | Best Performing Method | Performance Insight |
|---|---|---|
| Proteins with <25% Sequence Identity | Protein LLMs (ESM2) [1] | LLMs significantly outperform BLASTp, as they are not reliant on direct homology and can leverage learned biochemical patterns. |
| Specific EC Number Classes | Mixed Results [1] | The study found that BLASTp excels at predicting certain EC numbers, while LLMs are better at others, indicating their predictions are complementary. |
| Full-Annotation Difficulty | Protein LLMs (ESM2) [1] | ESM2 provides more accurate predictions on difficult annotation tasks where sequence clues are subtle or complex. |
Experimental Protocols: Methodology for Comparative Evaluation
To ensure the objectivity and reproducibility of the comparisons, the cited studies employ rigorous experimental frameworks. The following workflow and protocol detail describe a typical evaluation setup.
Diagram 1: Experimental Workflow for Comparative Performance Assessment
Detailed Experimental Methodology
The foundational study for this comparison employed the following protocol [1]:
Data Acquisition and Processing: Protein sequences and their EC numbers were extracted from the UniProtKB database (SwissProt and TrEMBL) in February 2023. To ensure data quality and avoid homology bias, only UniRef90 cluster representatives were retained. UniRef90 clusters group sequences that share at least 90% identity, with the representative chosen based on annotation quality and sequence length.
Problem Formulation: EC number prediction was framed as a global hierarchical multi-label classification problem. A single classifier is tasked with predicting the entire hierarchy of EC numbers (e.g., for EC 1.1.1.1, the model must correctly predict labels at levels 1, 1.1, 1.1.1, and 1.1.1.1). This accounts for promiscuous and multi-functional enzymes.
Model Training and Evaluation:
- BLASTp: For a given query sequence in the test set, BLASTp was run against a database of training sequences. The EC number of the top-hit (most similar sequence) was transferred to the query.
- Protein LLMs: Pre-trained models (ESM2, ESM1b, ProtBERT) were used as feature extractors. The embeddings (numerical representations) for each protein sequence were extracted and used to train a fully connected deep neural network (DNN) classifier. The performance of this DNN was then evaluated on the held-out test set.
Performance Metrics: Models were compared based on their accuracy in predicting the exact EC number on the test set. Further analysis segmented performance based on difficulty and sequence similarity to understand the specific scenarios where each method excels.
Successful implementation of these computational methods relies on key software tools and databases.
| Resource Name | Type | Primary Function in Research | Access / Reference |
|---|---|---|---|
| UniProtKB | Database | Provides the comprehensive, curated dataset of protein sequences and functional annotations (e.g., EC numbers) required for training and benchmarking. [1] [3] | https://www.uniprot.org/ |
| ESM2 | Protein LLM | A state-of-the-art evolutionary scale model used to convert protein sequences into numerical embeddings that capture contextual, functional information. [1] | https://github.com/facebookresearch/esm |
| BLASTp | Software Suite | The standard benchmark for sequence-similarity-based function prediction, used for comparative performance analysis. [1] | https://blast.ncbi.nlm.nih.gov/ |
| PyTorch / TensorFlow | Framework | Deep learning frameworks used to build and train the classifier neural networks on top of protein LLM embeddings. [1] | https://pytorch.org/, https://www.tensorflow.org/ |
| DIAMOND | Software Tool | A faster, more sensitive alternative to BLASTp for sequence alignment, often used in high-throughput pipelines. [1] | https://github.com/bbuchfink/diamond |
The evidence demonstrates that the choice between context-based Protein LLMs and sequence-based BLASTp is not a binary one. BLASTp maintains a slight overall edge in accuracy when reliable homologs are present, justifying its continued role in mainstream annotation pipelines. However, Protein LLMs, particularly ESM2, have proven superior for the critical task of annotating proteins with no or distant homologs, a common challenge in metagenomics and the study of under-characterized organisms [1].
The most powerful finding is that their predictions are complementary. Each method excels at predicting different subsets of EC numbers. Therefore, the future of high-accuracy automated protein function annotation lies not in selecting one over the other, but in their strategic integration. A hybrid pipeline that leverages the robust reliability of BLASTp for clear homologs and the powerful inference capabilities of Protein LLMs for difficult cases will provide the most comprehensive and accurate coverage, ultimately accelerating research in drug development and systems biology.
Conclusion
The competition between protein language models and BLASTp is not a zero-sum game; instead, the future of high-accuracy protein annotation lies in their strategic integration. While BLASTp remains the gold standard for routine annotation based on clear homology, pLMs have firmly established their superior capability for difficult cases, such as annotating enzymes with no close homologs or with very low sequence identity. The most powerful and effective annotation pipelines will therefore leverage the precision of BLASTp where applicable, while harnessing the pattern recognition and generalizability of advanced pLMs like ESM2 for the remaining challenging cases. For researchers in biomedicine and drug development, this synergistic approach promises to significantly accelerate the functional characterization of novel proteins, unlock new therapeutic targets, and drive innovation in protein engineering, ultimately closing the vast annotation gap in genomic databases.
References
- 1. Comparative Assessment of Protein Large Language ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11866580/]
- 2. Comparative Assessment of Protein Large Language ... [https://pubmed.ncbi.nlm.nih.gov/40016653/]
- 3. Evaluating the advancements in protein language models for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11790633/]
- 4. Faster, Better Results for Protein BLAST Searches [https://ncbiinsights.ncbi.nlm.nih.gov/2025/05/22/faster-better-results-protein-blast/]
- 5. Medium-sized protein language models perform well at ... [https://www.nature.com/articles/s41598-025-05674-x]
- 6. BASys2: a next-generation bacterial genome annotation ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12230659/]
- 7. From Sequence to Function with Protein Language Models [https://www.synbiobeta.com/read/from-sequence-to-function-with-protein-language-models]
- 8. A large-scale assessment of sequence database search tools ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11262835/]
- 9. Evaluating the advancements in protein language models ... [https://www.frontiersin.org/journals/bioengineering-and-biotechnology/articles/10.3389/fbioe.2025.1506508/full]
- 10. Comparative Assessment of Protein Large Language Models ... [https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-025-06081-9]
- 11. Sliding Window Interaction Grammar (SWING) [https://www.nature.com/articles/s41592-025-02723-1]
- 12. High-accuracy protein complex structure modeling based ... [https://www.nature.com/articles/s41467-025-65090-7]
- 13. [2510.11188] Protein as a Second Language for LLMs [https://arxiv.org/abs/2510.11188]
- 14. A Comprehensive Review of Protein Language Models [https://arxiv.org/html/2502.06881v1]
- 15. High attention sites as indicators of protein family and function ... [https://journals.plos.org/ploscompbiol/article?id=10.1371/journal.pcbi.1013424]
- 16. PLM-interact: extending protein language models to predict ... [https://www.nature.com/articles/s41467-025-64512-w]
- 17. Benchmarking protein language models for ... [https://www.nature.com/articles/s41598-025-86519-5]
- 18. Comparative assessment of annotation tools reveals ... [https://www.nature.com/articles/s41598-025-24333-9]
- 19. FANTASIA leverages language models to decode the ... [https://www.nature.com/articles/s42003-025-08651-2]
- 20. A High-Throughput Implementation of the ProtTrans Model [https://pubmed.ncbi.nlm.nih.gov/40601255/]
- 21. Annotating the genome at single-nucleotide resolution with ... [https://www.nature.com/articles/s41592-025-02881-2]
- 22. The PLSDB 2025 update: enhanced annotations and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11701622/]
- 23. BLAST: Basic Local Alignment Search Tool [https://blast.ncbi.nlm.nih.gov/Blast.cgi]
- 24. 2025 BLAST NEWS — BlastNews 0.1.1 documentation - NIH [https://blast.ncbi.nlm.nih.gov/doc/blast-news/2025-BLAST-News.html]
- 25. Top 10 Bioinformatics Tools in 2025: Features, Pros, Cons ... [https://www.scmgalaxy.com/tutorials/top-10-bioinformatics-tools-in-2025-features-pros-cons-comparison/]
- 26. Fine-tuning protein language models unlocks the potential of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12450373/]
- 27. PLMSearch: Protein language model powers accurate and ... [https://www.nature.com/articles/s41467-024-46808-5]
- 28. NCSP-PLM: An ensemble learning framework for ... [https://www.aimspress.com/article/doi/10.3934/mbe.2024063?viewType=HTML]
- 29. Autoregressive enzyme function prediction with multi-scale ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12448393/]
- 30. TopEC: prediction of Enzyme Commission classes by 3D ... [https://www.nature.com/articles/s41467-025-57324-5]
- 31. Researchers glimpse the inner workings of protein ... [https://news.mit.edu/2025/researchers-glimpse-inner-workings-protein-language-models-0818]
- 32. Research 4 [https://peterkimlab.stanford.edu/research-4]
- 33. Large language models generate functional protein ... [https://www.nature.com/articles/s41587-022-01618-2]
- 34. De novo protein design with a language model [https://www.nature.com/articles/s41587-022-01518-5]
- 35. Evaluating large language models for annotating proteins [https://pmc.ncbi.nlm.nih.gov/articles/PMC11070647/]
- 36. Multi-modal deep learning enables efficient and accurate ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11347633/]
- 37. Contrastive-learning of language embedding and ... [https://www.nature.com/articles/s41467-025-56526-1]
- 38. Benchmark Evaluation of Protein–Protein Interaction ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8746451/]
- 39. Prediction of protein–protein interaction based ... - BMC Biology [https://bmcbiol.biomedcentral.com/articles/10.1186/s12915-025-02359-9]
- 40. Protein BLAST: search protein databases using a protein query [https://blast.ncbi.nlm.nih.gov/Blast.cgi?PAGE=Proteins]
- 41. BLAST-based structural annotation of protein residues using ... [https://biologydirect.biomedcentral.com/articles/10.1186/s13062-016-0106-9]
- 42. Is Protein BLAST a thing of the past? [https://www.nature.com/articles/s41467-023-44082-5]
- 43. Are protein language models the new universal key? [https://www.sciencedirect.com/science/article/pii/S0959440X25000156]
- 44. identification of intrinsically disordered regions based on ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12231546/]
- 45. Benchmarking recent computational tools for DNA-binding ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11630855/]
- 46. [2510.23127] Lost in Tokenization: Context as the Key to ... [https://arxiv.org/abs/2510.23127]
- 47. pLM-BLAST: distant homology detection based on direct ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10576641/]
- 48. Biophysics-based protein language models for ... [https://www.nature.com/articles/s41592-025-02776-2]
- 49. T4SEpp: A pipeline integrating protein language models to ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10847861/]
- 50. Improving prediction performance of general protein ... [https://www.nature.com/articles/s41467-024-52293-7]
- 51. Accurately predicting enzyme functions through geometric ... [https://www.nature.com/articles/s41467-024-52533-w]
- 52. Protein language models: promises, pitfalls and applications [https://pipebio.com/blog/protein-language-models]
- 53. Review on the Computational Genome Annotation of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7565776/]
- 54. Accurate prediction of protein function using statistics- ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11297950/]